COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
7/6/2020
The Human Leukocyte Antigen (HLA) family – the name given to the Major Histocompatibility Complex in humans – is a highly polymorphic gene cluster involved in presentation of foreign antigens to the immune system and, importantly, in graft rejection (Choo, 2007). HLA molecules come in three main types (called, types I, II, and III); type I HLA are found on all nucleated cells in the body, and primarily present antigens derived from within the host cell, whether they be viral particles or host peptide fragments resulting from cell degradation. Type II HLA are found almost exclusively on the “professional” antigen-presenting cells: B cells, macrophages, and dendritic cells. These molecules present antigens that originate from outside host cells. Typically, these are fragments from extracellular bacteria and parasites, and environmental allergens. Finally, type III HLA encode accessory proteins: “non-classical” HLA molecules that aid in presentation, and components of the complement system.
The HLA system is considered to be one of the most polymorphic – that is, varied and diverse – gene families in the human genome. There are millions of variations of each type; and each individual carries six unique type I molecules (a I-A, I-B and I-C from mom, and a I-A, I-B, and I-C from dad). This means that the HLA system forms the molecular basis of “self”. Each person is genetically distinct in their specific combination of HLA molecules, even among type I alone. That said, the HLA family, like any other gene, is subject to variations in frequency in the population. And its association with immune function means that some of this variation is reflected in differential susceptibility to disease. For example, the HLA variant HLA-DQB1*02 is overrepresented in patients with celiac disease, and the type I HLA-B53 has been shown to protect against severe malaria (Choo, 2007; Hill et al., 1992).
Nguyen et al. performed an in silico analysis of the SARS-CoV-2 genome to determine which of the type I HLA alleles confer protection, and which enhance susceptibility. The authors first determined all possible peptides of lengths 8 to 12 amino acids that could be formed from the SARS-CoV-2 genome, for a total of 32,257 peptides. These peptides were tested against 145 different allelic variation of type I HLA molecules to assess the binding affinity of each HLA-peptide pair. They noted that the molecule HLA-B*46:01 had the fewest predicted binding peptides (the lowest binding affinity) of the 145 tested. When the same analysis was performed for the SARS-CoV-1 proteome, HLA-B*46:01 again had the lowest predicted binding affinity. They did not find the binding affinity across all 145 HLA alleles to be related to the global frequency distribution of a given allele (Figure 1). These data indicate that individuals who express HLA-B*46:01 may be more prone to severe COVID-19, due to the fact that their HLA system has a relatively low ability to present viral peptides to the immune system for initiation of an immune response.

In order to determine the extent of cross-reactive immunity that may be conferred by prior exposure to other human coronaviruses, the authors aligned reference sequence data for 5 essential coronavirus proteins – ORF1ab, spike (S), envelope (E), membrane (M), and nucleocapsid (N) – from 34 distinct coronaviruses. They identified 48 conserved amino acid sequences; 44 of these sequences they anticipated would produce at least one 8- to 12-mer peptide also found within at least one other human coronavirus (Figure 2), for a total of 564 peptides. The binding affinity analysis was repeated for these potentially cross-reactive peptides. It was demonstrated that the alleles HLA-A*02:02, HLA-B*15:03, and HLA-C*12:03 had the highest predicted binding affinity for conserved peptides. However, 56 different alleles (including HLA-B*46:01) demonstrated no significant binding affinity to any conserved peptides. The authors note that this suggests a lack of potential cross-protective immunity in individuals expressing any of these 56 alleles. Again, they indicate that capacity for conserved peptide presentation among the 564 tested alleles is not correlated with the frequency of the given alleles in the global population (Figure 3).


In all, these data indicate that at least some level of protection, however insignificant, may result from the random allotment of an individual’s HLA type I alleles. These data also indicate that individuals expressing HLA alleles on the lower end of the protection spectrum may actually have a heightened susceptibility to infection and severity of disease, relative to other individuals, based on their inability to present viral peptide antigens to the immune system. Via genetic testing, this information may be used to identify individuals at a higher risk of contracting the disease, and on a larger scope, may be used to identify patterns of heightened susceptibility or protection at the population level. The authors of this paper note the inherent limitations of in silico analyses – it is important to remember that the data and figures presented here are largely predictions, however accurate they may ultimately be. Therefore, future work should focus on performing real-world genetic analyses of COVID-19 patients in order to determine the correlations between genotype and severity of contracted disease.
References
Choo, S. Y. (2007). The HLA system: Genetics, immunology, clinical testing, and clinical implications. Yonsei Medical Journal, 48(1), 11–23. https://doi.org/10.3349/ymj.2007.48.1.11
Hill, A. V. S., Elvin, J., Willis, A. C., Aidoo, M., Allsopp, C. E. M., Gotch, F. M., Ming Gao, X., Takiguchis, M., Greenwood, B. M., Townsend, A. R. M., McMichael, A. J., & Whittle, H. C. (1992). Molecular analysis of the association of HLA-B53 and resistance to severe malaria. Nature, 360(6403), 434–439. https://doi.org/10.1038/360434a0
Nguyen, A., David, J. K., Maden, S. K., Wood, M. A., Weeder, B. R., Nellore, A., & Thompson, R. F. (2020). Human leukocyte antigen susceptibility map for SARS-CoV-2. Journal of Virology, 94(13). https://doi.org/10.1128/jvi.00510-20
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
6/30/2020
Much of the research thus far on the host immune response to SARS-CoV-2 infection has been geared toward the development of novel vaccines, and for this reason has focused largely on the roles of T and B cells in fighting infection. However, another class of immune cells, the so-called “natural killer” cells, are central to the response to viral infection as well. Natural killer (NK) cells are lymphocytes that are derived from the same progenitor cell line as T and B cells but are non-specific in effect and therefore part of the innate immune branch (Trinchieri, 1989). These cells play a large part in the recognition and elimination of virally infected cells and tumor cells, in which the Major Histocompatibility Complex class I (MHC I) molecules are often downregulated or mutated in an “effort” by the hijacked cell to avoid initiating an immune response. In the absence of MHC I on the surface of a host cell, the inhibition receptors on NK cells cannot be cross-linked, and the lymphocytes are subsequently activated against the pathological host cell(s) (Caligiuri, 2008; Long et al., 2013). The role of NK cells in fending against SARS-CoV-2 cannot be overstated and determining the virus’ interaction with these cells in vivo is an important next step in understanding the complete host immune response.
Bortolotti et al. investigated the interaction between the coronavirus spike proteins and NK cells in context of exhaustion of these lymphocytes. Previous work this year has demonstrated significant functional exhaustion of NK cells and CD8+ T cells in COVID-19 patients, mediated by the upregulation of the inhibitory receptor CD94/NKG2A (Zheng et al., 2020). In this study, peripheral blood natural killer cells were harvested from healthy control patients who tested negative for SARS-CoV-2, thus ensuring that these cells were entirely naïve for the spike proteins of the virus. These NK cells were cultured in the presence of either the SARS-CoV-2 spike protein domain 1 (SP1), SARS-CoV-2 spike protein domain 2 (SP2), or SARS-CoV spike protein (S), and their migration along a polycarbonate filter was measured. The authors determined that all three spike protein treatments induced significant migration compared to the control chemoattractant (CXCL12), confirming the ability of the spike protein to induce NK cell trafficking to sites of infection (Figure 1A). As a follow up, interferon-gamma (IFN-γ) expression was observed in the presence of each spike protein as a measure of NK cell activation: indeed, an increase in IFN-γ was noted in the presence of SP1, but not SP2 or S protein (Figure 1B).

The authors then tested the effects of spike protein on the cytotoxicity of NK cells. To do so, SP1, SP2, and S protein were expressed in K562 cells, and the harvested NK cells were incubated with the K562 cells, after which expression of the molecule CD107a (a marker of NK cell degranulation) was observed. At the end of the incubation period, they noted a non-significant increase in CD107a expression for each of the three treatments compared to control K562 cells (Figure 2, top). Due to the respiratory nature of the virus, however, the experiment was repeated in the lung epithelial cell line Beas-2B. This time, the authors observed a significant decrease in CD107a in the cells expressing SP1; and a non-significant decrease in CD107a was observed in the SP2- and S-expressing cells (Figure 2, bottom). They concluded that impairment of CD107a expression indicates a reduction in the cytotoxic capacity of the NK cells due to interaction with SP1.

Finally, to determine the exact mechanism of the reduced cytotoxicity, the authors expressed SP1, SP2, and S in Beas-2B cells and stained them with anti-classical Human Leukocyte Antigen I antibodies (anti-HLA-A, anti-HLA-B, and anti-HLA-C). They observed a significant decrease in HLA I cell surface expression in Beas-2B cells that were transfected with SP1, indicating that SP1 has a large role in inhibiting the presentation of viral antigens by the host cells (Figure 3, top; see updates 6/15/2020 for discussion of MHC [HLA] I presentation). HLA-E is a “non-classical” HLA I molecule that is expressed on the surfaces of many cell types and interacts specifically with the CD94/NKG2A inhibitory receptors on NK cells. When the researchers stained the transfected Beas-2B cells with anti-HLA-E antibody, they noted a significant increase in HLA-E surface expression in the presence of SP1, but not SP2 or S (Figure 3, middle). Lastly, when NK cells were incubated with SP1-transfected Beas-2B cells, there was a significant increase in CD94/NKG2A expression on the NK cell surfaces (Figure 3, bottom). Taken together, these data indicate the ability of the SARS-CoV-2 SP1 to significantly downregulate NK cell activity in vivo via a heightened interaction between HLA-E and the inhibitory receptor CD94/NKG2A.
One limitation of this study is the fact that the subunits of the SARS-CoV-2 spike protein were only tested individually. It is not clear why the SP1 subunit should exert the observed effects on NK cells when the SP2 subunit does not. Further characterization of the function(s) of each subunit may shed light on the reason behind this disparity. It is not likely that the SP1 subunit is viable to initiate and sustain infection on its own, and it is likely not the case that the subunits of the spike protein would be encountered individually anyway. Future work should focus on determining the effects of the entire SARS-CoV-2 spike protein on NK cell cytotoxicity and exhaustion.

References
Bortolotti, D., Gentili, V., Rizzo, S., & Rotola, A. (2020). Sars-Cov-2 Spike 1 protein control Natural killer cells activation via HLA-E / NKG2A pathway. 1–18. https://doi.org/10.21203/RS.3.RS-31860/V1
Caligiuri, M. A. (2008). Human natural killer cells. Blood, 112(3), 461–469. https://doi.org/10.1182/blood-2007-09-077438
Long, E. O., Sik Kim, H., Liu, D., Peterson, M. E., & Rajagopalan, S. (2013). Controlling Natural Killer Cell Responses: Integration of Signals for Activation and Inhibition. Annual Review of Immunology, 31(1), 227–258. https://doi.org/10.1146/annurev-immunol-020711-075005
Trinchieri, G. (1989). Biology of Natural Killer Cells. Advances in Immunology, 47(C), 187–376. https://doi.org/10.1016/S0065-2776(08)60664-1
Zeng, F., Dai, C., Cai, P., Wang, J., Xu, L., Li, J., Hu, G., Wang, Z., Zheng, F., & Wang, L. (2020). A comparison study of SARS-CoV-2 IgG antibody between male and female COVID-19 patients: A possible reason underlying different outcome between sex. Journal of Medical Virology, jmv.25989. https://doi.org/10.1002/jmv.25989
Zheng, M., Gao, Y., Wang, G., Song, G., Liu, S., Sun, D., Xu, Y., & Tian, Z. (2020). Functional exhaustion of antiviral lymphocytes in COVID-19 patients. In Cellular and Molecular Immunology (Vol. 17, Issue 5, pp. 533–535). Springer Nature. https://doi.org/10.1038/s41423-020-0402-2
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
6/15/2020
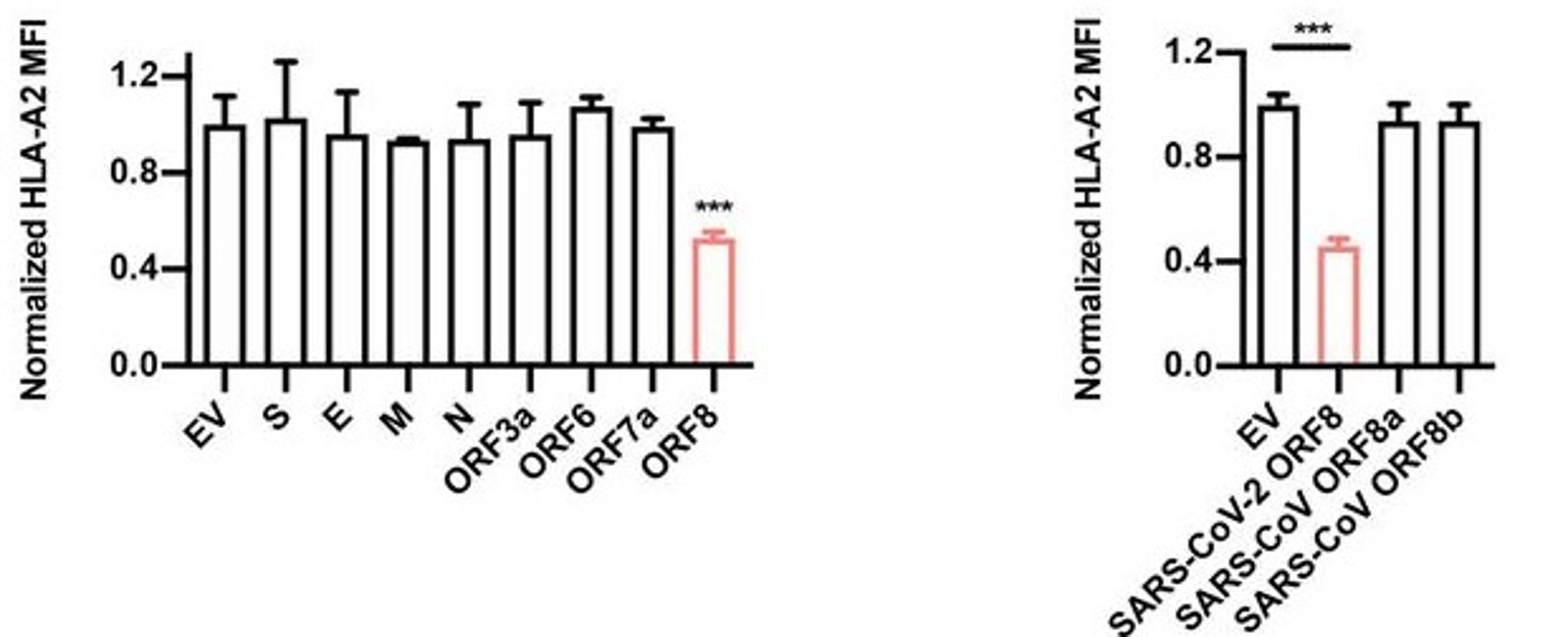
Though much progress has been made in determining the specific interactions between SARS-CoV-2 infectious particles and host cells (Liang et al., 2020; see updates for week of 6/2/2020), we are only beginning to understand the exact mechanisms of pathogenicity in detail. As part of this ongoing inquiry, Zhang et al. surveyed the four structural proteins of SARS-CoV-2 (spike [S], envelope [E], membrane [M], and nucleocapsid [N]) and several accessory proteins found throughout the virus’ genome to assess what effects they have on pathogenicity. The authors found that one accessory protein, encoded by open reading frame (ORF) 8, was able to significantly inhibit a host cell’s ability to initiate an immune response by downregulating expression of the Major Histocompatibility Complex (MHC) I protein by these cells (Figure 1).
Briefly, the MHC is a family of molecules that comes in two major classes (I and II), each composed of multiple subclasses and allotypes that give rise to a diverse array of functional proteins. Both MHC I and II are involved with the presentation of antigens and activation of immune responses, but they act at separate, yet interdependent, stages. MHC I is a molecule that is expressed on almost all of a host organism’s cells (non-nucleated cells like red blood cells do not express MHC I), and allows presentation of antigens to innate immune cells – typically dendritic cells – that then relay the signal to T cells in lymph nodes and the spleen (York, Rock, 1996). The presence of functional MHC I on the surface of host cells is required for proper development of CD8+ T cells, and therefore for the development of a potent cell-mediated immune response. Any manner of interference that alters the components of the molecule or its associated protein complexes (such as tapasin, the gatekeepers TAP1 and TAP2, or an enzyme called ERp57, among the proteins that make up the peptide loading complex of MHC I; Blees et al., 2017) will inhibit proper functioning. Similarly, mutations or interactions with pathogenic molecules may prevent MHC I from binding an antigen within the cell and being trafficked to the surface, thereby acting to suppress the immune response of the organism and diminish its capacity to clear an infection. This interference may manifest in the form of random mutations, or via interactions with non-host molecules, as in the case of viral infection.
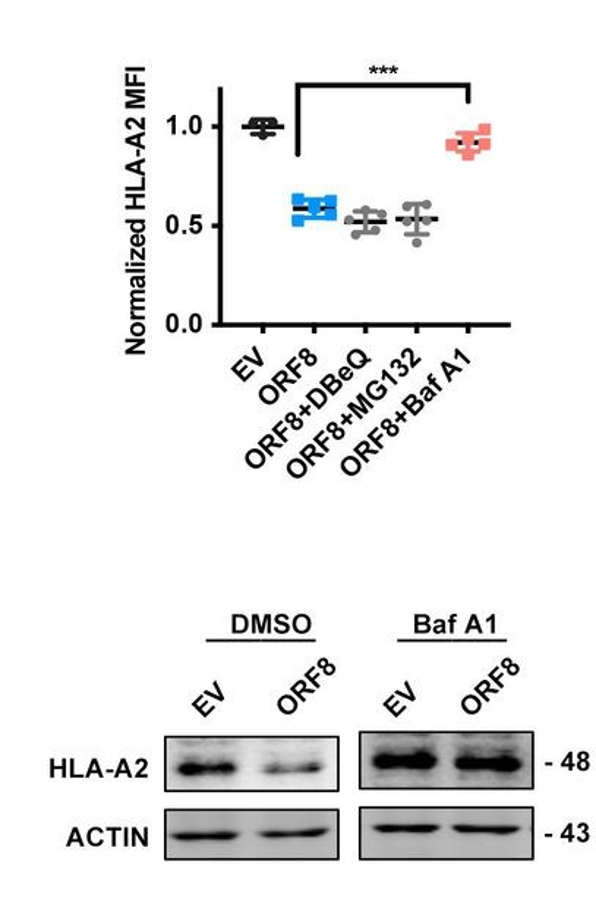
By transfecting HEK293T cells with ORF8-expressing or control-expressing plasmids, the authors of this study were able to determine in great detail the mechanism of the observed downregulation of MHC I. Using flow cytometry, they found that both the frequency and the mean fluorescence intensity of the heavy chain and beta-2-microglobulin components of the intact MHC I protein were significantly reduced in the ORF8-expressing cells relative to the control cells (Figure 2, top row), and that this reduction occurred in a dose-dependent manner. They then repeated the same plasmid experiment using other cell lines (human fetal colon cells, human bronchial epithelial cells, and human liver cells) and, in a separate experiment, infected HEK293T with authentic SARS-CoV-2 viral particles, all of which yielded results similar to the original trial – in other words, MHC I was downregulated regardless of cell line or origin of the expressed ORF8 protein (Figure 2, bottom row). When HEK cells were treated with various inhibitors of protein degradation, only bafilomycin A1, an inhibitor of lysosomal degradation, significantly counteracted the effects of ORF8 (Figure 3). The other inhibitors tested (N2, N4-dibenzylquinazoline-2,4-diamine [DBeQ]; and MG132) block endoplasmic reticulum-associated protein degradation and ubiquitin-protease-associated degradation, respectively. These results suggest a role for ORF8 in the targeting of MHC I complex molecules to lysosomes for degradation. Such sequestration ensures that the MHC molecules cannot bind antigen within endosomal compartments and never reach the cell surface for presentation of this antigen. This interaction is immunosuppressive for the reasons outlined above.
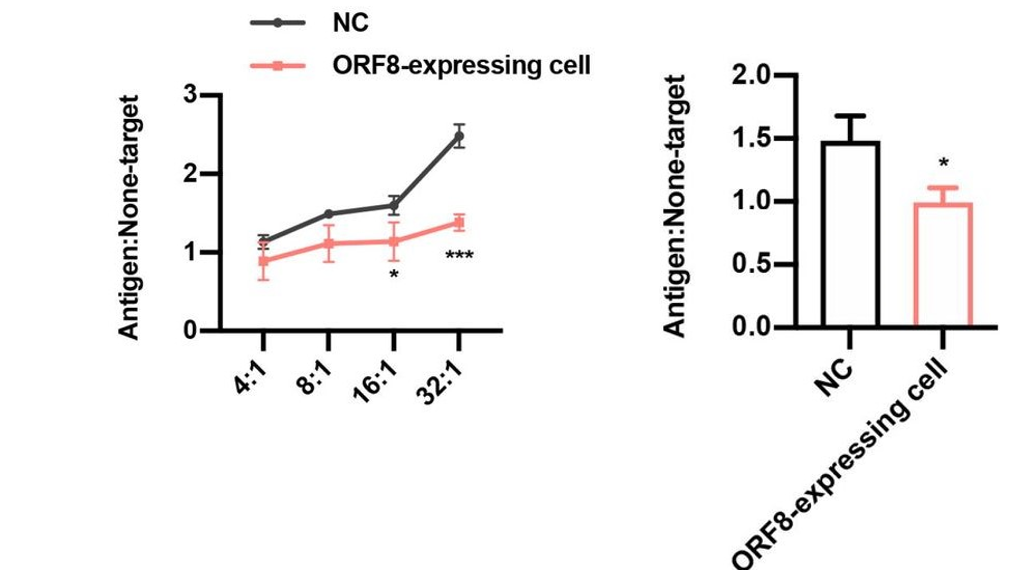
SARS-CoV spike protein-derived peptide-1 (SSp-1) is an epitope (the specific region of an antigen on the scale of molecules or even atoms that is recognized by immune cells) present in the spike protein of both SARS-CoV and SARS-CoV-2. The overall sequence homology of the spike protein between these two viruses is relatively low. However, this specific epitope represents an island of similarity that is hypothesized as a potential target through which the host immune system may recognize SARS-CoV-2, given its involvement in the recognition of SARS-CoV (Grifoni et al., 2020). When SSp-1-specific cytotoxic lymphocytes (CTLs) were directed against ORF8-expressing and control HEK293T cells, the lymphocytes were only able to eliminate the ORF8-expressing cells with a much lower efficiency than the control cells (Figure 4, left). And when CD8+T cells from recovered COVID-19 patients were directed against the same sets of HEK cells, they too could only eliminate the ORF8-expressing cells with an impaired efficiency relative to the control cells (Figure 4, right). These results reflect the inability of the HEK cells to successfully express antigen on their surface, due to a lack of functional MHC I protein.
In all, this paper marks an important step toward understanding the exact mechanisms of SARS-CoV-2 pathogenicity. Developing an understanding of specific interactions between SARS-CoV-2 molecules and host molecules (for example, the interaction between viral non-structural proteins 9/10 and host nuclear factor-kappa B-repressing protein [Liang et al., 2020], or between ORF8 and MHC I outlined here) will move us further in the direction of producing therapies that are incredibly potent and, importantly, virus specific. The authors note that their findings suggest a mechanism for autophagy mediated by the endoplasmic reticulum that is responsible for the transference of MHC I to lysosomes. The elucidation of this mechanism, if it indeed exists, should be a priority for the continuation of the work outlined here. Future studies should focus on understanding the specifics of these processes and on developing therapies that exploit faults in these interactions to fight infection.

References
Blees, A., Januliene, D., Hofmann, T., Koller, N., Schmidt, C., Trowitzsch, S., Moeller, A., & Tampé, R. (2017). Structure of the human MHC-I peptide-loading complex. Nature, 551(7681), 525–528. https://doi.org/10.1038/nature24627
Grifoni, A., Sidney, J., Zhang, Y., Scheuermann, R. H., Peters, B., & Sette, A. (2020). Candidate Targets for Immune Responses to 2019-Novel Coronavirus (nCoV): Sequence Homology- and Bioinformatic-Based Predictions. SSRN Electronic Journal. https://doi.org/10.2139/ssrn.3541361
Thieme, C. J., Anft, M., Paniskaki, K., Blazquez-Navarro, A., Doevelaar, A., Seibert, F. S., Hoelzer, B., Konik, M. J., Brenner, T., Tempfer, C., Watzl, C., Dolff, S., Dittmer, U., Westhoff, T. H., Witzke, O., Stervbo, U., Roch, T., & Babel, N. (2020). The SARS-CoV-2 T-cell immunity is directed against the spike, membrane, and nucleocapsid protein and associated with COVID 19 severity. MedRxiv, 2020.05.13.20100636. https://doi.org/10.1101/2020.05.13.20100636
York, I. A., & Rock, K. L. (1996). Antigen Processing and Presentation by the Class I Major Histocompatibility Complex. Annual Review of Immunology, 14(1), 369–396. https://doi.org/10.1146/annurev.immunol.14.1.369
Zhang, Y., Zhang, J., Chen, Y., Luo, B., Yuan, Y., Huang, F., Yang, T., Yu, F., Liu, J., Liu, B., Song, Z., Chen, J., Pan, T., Zhang, X., Li, Y., Li, R., Huang, W., Xiao, F., & Zhang, H. (2020). The ORF8 Protein of SARS-CoV-2 Mediates Immune Evasion through Potently Downregulating MHC-I. BioRxiv. https://doi.org/10.1146/annurev.immunol.14.1.369
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
6/8/2020
In the process of lymphocyte development, progenitor T and B cells undergo several rounds of signaling and selection before being released to the periphery to fight infections. As part of the normal developmental routine, immature lymphocytes undergo “negative selection” in which cells bearing autoreactive (self-reactive) antigen receptors are removed from the population, thereby preventing potential future reactions against the host’s own body (Palmer, 2003; Melchers et al., 1995).
Viral infections often cause major damage to host cells and tissues, setting the stage for serious and even life-threatening secondary sequelae. In particular, autoimmune conditions can develop as a result of viral infections via a number of mechanisms: a dysregulated inflammatory response can continue damaging the host even after the pathogen has been cleared; infection of “immunologically privileged” sites – tissues generally sequestered from the central and peripheral immune systems like the brain, eyes, germ cells, etc. – can cause the release from these tissues of antigens to which the peripheral immune system was previously ignorant, leading to an autoimmune response; and viral proteins can even retain enough similarity to host proteins to elicit the formation of antibodies that, consequently, are also autoreactive. These mechanisms present a “workaround” for the development of autoimmunity despite the successful (though usually incomplete) purging of autoreactive lymphocytes in normal development.
Immune thrombocytopenic purpura (ITP) is one such autoimmune condition that has been established as a sequela of pathogenic infection. Specifically, its development and progression has been observed post-infection with the human immunodeficiency virus (HIV)-1, hepatitis C virus (HCV), the bacteria Helicobacter pylori, and even SARS-CoV-1 (Liebman, 2008; Cines, Blanchette, 2002). ITP is characterized by a low platelet count (thrombocytopenia) and mucocutaneous bleeding and comes in two major forms, primary (idiopathic) and secondary, which have slightly different presentations and courses of progression. In primary ITP, autoreactive IgG attacks the body’s own platelets and opsonizes them for clearance in the spleen and liver – that is, the IgG molecules coat the surface of the platelets and mark them for destruction by innate leukocytes like macrophages (Cines, Blanchette, 2002). Secondary ITP is just that: it occurs secondarily to some other underlying condition, whether that is a lymphoproliferative disorder like chronic lymphoblastic leukemia or Hodgkin’s lymphoma, or an infection. Secondary ITP is characterized by accelerated platelet destruction and/or impaired platelet production, though the source of this pathology can vary from patient to patient. Treatment of secondary ITP generally focuses on targeting the underlying condition rather than on restoring platelet counts directly (Liebman, Stasi, 2007; Liebman, 2009).
Recently, ITP has been diagnosed in several COVID-19 patients. Zulfiqar et al. reported a case of a 65-year-old woman who presented to the emergency department with hypertension and autoimmune hypothyroidism, fatigue, fever, dry cough, and abdominal discomfort. Upon examination, she had a normal white blood cell count, normal hemoglobin, and normal platelet counts, but a slightly elevated C-reactive protein (CRP) level, indicative of an ongoing inflammatory response. Indeed, she subsequently tested positive for COVID-19 via oropharyngeal swab. On day 4 post-admission, the patient developed purpura on her lower extremities (Figure 1) and had a platelet count of 66,000 platelets/mm3, which dropped to 8,000/mm3 by day 7 (normal platelet counts are between ~150,000/mm3 and ~450,000/mm3). Two doses of intravenous immunoglobulin (IVIg) had little effect on the patient’s thrombocytopenia; a platelet transfusion on day 9 combined with eltrombopag (75 mg/day) led to recovered platelet counts (139,000/mm3) by day 13, at which time the purpura also disappeared (Zulfiqar et al., 2020).
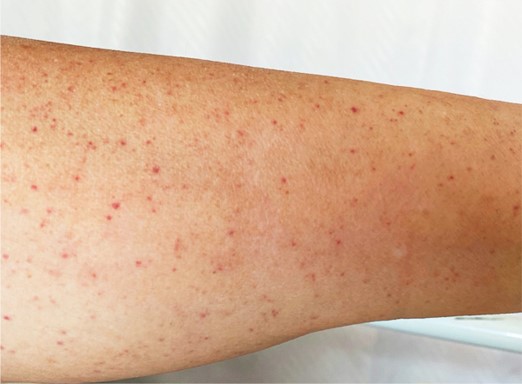
Another case of ITP following SARS-CoV-2 infection was reported recently in the Journal of Medical Cases. An 86-year-old female presented to the hospital with dyspnea (trouble breathing) and a cough, as well as petechial lesions on her limbs, lower back, and oral mucosa that had appeared 24 hours prior to her admission (Figure 2). No abnormal leukocytosis was observed, though the patient did have a moderate CRP elevation, again indicative of an inflammatory response. Her complete blood count (CBC) demonstrated severe thrombocytopenia (7000/mm3), and she tested PCR-positive for SARS-CoV-2 infection. Platelet transfusions occurred on day 1 and day 2 post-admission, though without significant initial effect. Corticosteroid treatment, often used to ameliorate severe inflammation, was started on day 2 in addition to IVIg, and by day 7 the platelet count had risen to 144,000/mm3. The patient was discharged two weeks later with a stable platelet count (Gonze et al., 2020).

Thrombocytopenia is a condition that was frequent as a result of SARS-CoV-1 infection, but ITP, though associated with other viruses, is relatively rare in association with coronaviruses. In the original SARS coronavirus outbreak, ITP was found to be triggered by various antibiotics and antivirals, heparin, hemodialysis, and even extracorporeal membrane oxygenation (ECMO). Though currently unknown, the mechanisms of thrombocytopenia, both immune and non-immune, in COVID-19 are hypothesized to be similar to the mechanisms observed in SARS-CoV-1 infection: decreased platelet production caused by the cytokine storm, and infection of bone marrow stromal cells and hematopoietic stem cells; increased platelet destruction as a result of autoreactive antibodies and the formation of immune complexes; and a decreased circulating platelet count due to lung injury (Yang et al., 2020). The fact that steroids were effective in restoring platelet counts in these few case studies may suggest the significant role played by the inflammatory cytokine storm in inducing thrombocytopenia, as these steroids would have helped moderate the effects of the cytokine release. Again, ITP is relatively rare in SARS-CoV-2 infection, but acts as a marker of potential “side effects” of infection and presents a new angle to take into consideration when determining the best course of treatment for a given patient.
References
Cines, D. B., & Blanchette, V. S. (2002). Medical progress: Immune thrombocytopenic purpura. In New England Journal of Medicine (Vol. 346, Issue 13, pp. 995–1008). Massachusetts Medical Society. https://doi.org/10.1056/NEJMra010501
Gonze, A., Gonze, A., Hannesse, C., Coppens, N., Vellemans, H., & Maury, G. (2020). SARS-CoV-2-Induced Severe Immune Thrombocytopenic Purpura. Journal of Medical Cases, 11(6), 166–168. https://doi.org/10.14740/jmc.v11i6.3481
Liebman, H. A. (2008). Viral-associated immune thrombocytopenic purpura. Hematology / the Education Program of the American Society of Hematology. American Society of Hematology. Education Program, 2008(1), 212–218. https://doi.org/10.1182/asheducation-2008.1.212
Liebman, H. A. (2009). Recognizing and Treating Secondary Immune Thrombocytopenic Purpura Associated With Lymphoproliferative Disorders. Seminars in Hematology, 46(SUPPL. 2). https://doi.org/10.1053/j.seminhematol.2008.12.004
Liebman, H. A., & Stasi, R. (2007). Secondary immune thrombocytopenic purpura. In Current Opinion in Hematology (Vol. 14, Issue 5, pp. 557–573). https://doi.org/10.1097/MOH.0b013e3282ab9904
Melchers, F., Rolink, A., Grawunder, U., Winkler, T. H., Karasuyama, H., Ghia, P., & Andersson, J. (1995). Positive and negative selection events during B lymphopoiesis. Current Opinion in Immunology, 7(2), 214–227. https://doi.org/10.1016/0952-7915(95)80006-9
Palmer, E. (2003). Negative selection – Clearing out the bad apples from the T-cell repertoire. In Nature Reviews Immunology (Vol. 3, Issue 5, pp. 383–391). Nature Publishing Group. https://doi.org/10.1038/nri1085\
Yang, Y., Zhao, J., Wu, J., Teng, Y., & Xia, X. (2020). A Rare Case of Immune Thrombocytopenic Purpura Secondary to COVID‐19. Journal of Medical Virology, jmv.26051. https://doi.org/10.1002/jmv.26051
Zulfiqar, A. A., Lorenzo-Villalba, N., Hassler, P., & Andrès, E. (2020). Immune Thrombocytopenic Purpura in a Patient with COVID-19. In New England Journal of Medicine (Vol. 382, Issue 18, p. E43). Massachussetts Medical Society. https://doi.org/10.1056/NEJMc2010472
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
6/2/2020
The role of interferons in fending against viruses has been known and investigated for decades, and in light of this, interferon therapy is well-established for treating viral infections and even some cancers (Sen, 2001; Samuel, 2001; Korenman et al., 1991; Tanner et al., 1990). Interferon therapy has recently been suggested as an option for use in COVID-19 treatment to mitigate the cytokine storm symptoms observed in early infection (Sallard et al., 2020; Prompetchara, Ketloy, Palaga, 2020). Less is known, however, about the exact molecular mechanisms that occur downstream due to interferon release.
Lymphocyte antigen 6 complex, locus E (LY6E) is one such molecule involved in the downstream action of interferon release. It is one protein in a class known as Interferon-Stimulated Genes (ISGs) and plays a central role in various immune and tumorigenic pathways (Xu et al., 2014; Yeom et al., 2016; Alhossiny et al., 2016). Pfaender et al. developed a study to determine the involvement of LY6E in SARS-CoV-2 infection. They screened over 350 previously known ISG DNA sequences for ability to inhibit the human coronavirus strain HCoV-229E; LY6E stood out as a potent inhibitor of infection. LY6E also significantly inhibited infection by HCoV-OC43, MERS-CoV, SARS-CoV, and SARS-CoV-2, in addition to hepatitis virus and murine coronavirus (Figure 1). To determine the effect that endogenous LY6E has in controlling infection, the CRISPR-Cas9 system was used to ablate LY6E in human lung adenocarcinoma cells, which exhibited significant HCoV-229E infection compared to un-ablated control cells. Subsequent reconstitution with CRISPR-resistant LY6E restored the antiviral activity in the knockout cells. Several mechanisms of antiviral action were tested for LY6E: it did not exhibit a capacity to restrict HCoV-229E attachment to host cells, nor did it demonstrate any effect on the expression of CD13, the host cell surface receptor for HCoV-229E. In a syncytia formation assay, however, vesicular stomatitis virus (VSV) pseudoparticles were made to express spike (S) proteins from each of the MERS, SARS, and SARS-2 coronaviruses and LY6E significantly inhibited syncytia formation by blocking membrane fusion (Pfaender et al., 2020). These data underscore the effect of interferon therapies for preventing viral infection. Future work should focus on elucidating ISG pathways, which may be key in developing more effective therapies, tailored to a given disease.


Viruses are not technically living beings and are characterized as “obligate intracellular” pathogens, meaning that they rely on their host cell’s machinery to replicate and propagate infections. It follows, then, that virus- and host-derived molecules mingle inside the infected cell, sometimes with the effect of modulating the activity of the molecule with which they interact. Liang et al. analyzed the network between SARS-CoV-2 and HEK293 cell proteins and developed a map of 251 animal (HEK) proteins that coordinate with the ~30 known coronavirus proteins to produce upwards of 630 high-confidence interactions (Figure 3). The authors noted several specific protein-protein synergies that they hypothesized to be involved in SARS-CoV-2 pathogenesis: between animal C1QBP and CoV nonstructural proteins [nsp] 9 and 10; between animal RAE1 and CoV open reading frame [ORF] 6; and between animal nuclear factor-kappa B-repressing protein [NKRF] and CoV nsp9, nsp10, nsp12, nsp13, and nsp15.
To determine which of these may be significant, peripheral blood mononuclear cells (PBMCs) from 35 COVID-19 patients were analyzed and revealed a heightened innate response compared to healthy controls, and especially elevations in interleukin (IL) 8 and IL6 that were significant in severe patients compared to mild patients and controls (Figure 4). In normal healthy humans, nuclear factor-kappa B (NF-κB) acts as a transcriptional hub to regulate the expression of many genes involved in the immune response and other biological systems (Bender et al., 2020; Meng et al., 2020; Tripathi, Aggarwal, 2006). Among these complex and interwoven pathways, it serves to induce the expression of IL8 and IL6, molecules critical to the production of inflammation in the early stages of the innate immune response (Kunsch, Rosen, 1993). NKRF is a molecule whose basic function is in the infancy of understanding, but which seems to be especially involved in inhibiting the action of NF-κB by degrading its downstream RNA products (Coccia et al., 2017; Memet et al., 2017). The elevated IL8/6 in severe COVID-19 patients and the observed interaction of NKRF with the coronavirus nonstructural proteins led the authors to hypothesize that nsp9/10/12/13/15 associate with NKRF during infection, thereby sequestering it and preventing it from inhibiting the induction of IL8 and IL6 by NF-κB.
Indeed, individually expressed nsp9 and nsp10 facilitated the significant induction of IL8 and IL6 in lung epithelial cells compared to controls, whereas nsp12, nsp13, and nsp15 did not (Figure 5). In addition, degradation of NKRF by silencing RNA in vitro resulted in an induction of IL8 similar to that seen in severe COVID-19 patients, thereby providing support for the authors’ hypothesis (Liang et al., 2020).



References
Alhossiny, M., Luo, L., Frazier, W. R., Steiner, N., Gusev, Y., Kallakury, B., Glasgow, E., Creswell, K., Madhavan, S., Kumar, R., & Upadhyay, G. (2016). Ly6E/K signaling to TGFβ promotes breast cancer progression, immune escape, and drug resistance. Cancer Research, 76(11), 3376–3386. https://doi.org/10.1158/0008-5472.CAN-15-2654
Bender, A. T., Tzvetkov, E., Pereira, A., Kasar, S., Przetak, M., Okitsu, S., & Vlach, J. (2020). TLR7 and TLR8 differentially activate the IRF and NF-kB pathways in specific cell types to promote inflammation. Lupus Science & Medicine, 7(Suppl 1), A74.1-A74. https://doi.org/10.1136/lupus-2020-eurolupus.138
Coccia, M., Rossi, A., Riccio, A., Trotta, E., & Santoro, M. G. (2017). Human NF-κB repressing factor acts as a stress-regulated switch for ribosomal RNA processing and nucleolar homeostasis surveillance. Proceedings of the National Academy of Sciences of the United States of America, 114(5), 1045–1050. https://doi.org/10.1073/pnas.1616112114
Korenman, J., Baker, B., Waggoner, J., Everhart, J. E., Di Bisceglie, A. M., & Hoofnagle, J. H. (1991). Long-term remission of chronic hepatitis B after alpha-interferon therapy. Annals of Internal Medicine, 114(8), 629–634. https://doi.org/10.7326/0003-4819-114-8-629
Kunsch, C., & Rosen, C. A. (1993). NF-kappa B subunit-specific regulation of the interleukin-8 promoter. Molecular and Cellular Biology, 13(10), 6137–6146. https://doi.org/10.1128/mcb.13.10.613
Liang, Q., Li, J., Guo, M., Tian, X., Liu, C., Wang, X., Yang, X., Wu, P., Xiao, Z., Qu, Y., Yin, Y., Fu, J., Zhu, Z., Liu, Z., Peng, C., & Zhu, T. (2020). Virus-host interactome and proteomic survey of PMBCs from COVID-19 patients reveal potential virulence factors influencing SARS-CoV-2 pathogenesis. BioRxiv, 2020.03.31.019216. https://doi.org/10.1101/2020.03.31.019216
Memet, I., Doebele, C., Sloan, K.E., Bohnsack, M.T. (2017). The G-patch protein NF-κB-repressing factor mediates the recruitment of the exonuclease XRN2 and activation of the RNA helicase DHX15 in human ribosome biogenesis. Nucleic Acids Research, 45(9), 5359-5374. https://doi.org/10.1101/2020.03.31.019216
Meng, Q., Liang, C., Hua, J., Zhang, B., Liu, J., Zhang, Y., Wei, M., Yu, X., Xu, J., & Shi, S. (2020). A miR-146a-5p/TRAF6/NF-kB p65 axis regulates pancreatic cancer chemoresistance: Functional validation and clinical significance. Theranostics, 10(9), 3967–3979. https://doi.org/10.7150/thno.40566
Pfaender, S., Mar, K. B., Michailidis, E., Kratzel, A., Hirt, D., V’kovski, P., Fan, W., Ebert, N., Stalder, H., Kleine-Weber, H., Hoffmann, M., Hoffmann, H. H., Saeed, M., Dijkman, R., Steinmann, E., Wight-Carter, M., Hanners, N. W., Pöhlmann, S., Gallagher, T., … Thiel, V. (2020). LY6E impairs coronavirus fusion and confers immune control of viral disease. BioRxiv, 2020.03.05.979260. https://doi.org/10.1101/2020.03.05.979260
Prompetchara, E., Ketloy, C., & Palaga, T. (2020). Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. In Asian Pacific Journal of Allergy and Immunology (Vol. 38, Issue 1, pp. 1–9). https://doi.org/10.12932/AP-200220-0772
Sallard, E., Lescure, F. X., Yazdanpanah, Y., Mentre, F., & Peiffer-Smadja, N. (2020). Type 1 interferons as a potential treatment against COVID-19. Antiviral Research, 178, 104791. https://doi.org/10.1016/j.antiviral.2020.104791
Samuel, C. E. (2001). Antiviral actions of interferons. In Clinical Microbiology Reviews (Vol. 14, Issue 4, pp. 778–809). American Society for Microbiology Journals. https://doi.org/10.1128/CMR.14.4.778-809.2001
Sen, G. C. (2001). Viruses and Interferons. Annual Review of Microbiology, 55(1), 255–281. https://doi.org/10.1146/annurev.micro.55.1.255
Tanner, D. J., Taylor, E. L., Smiles, K. A., & Peets, E. A. (1990). Intralesional interferon therapy for basal cell carcinoma. Journal of the American Academy of Dermatology, 23(4), 694–700. https://doi.org/10.1016/0190-9622(90)70276-N
Tripathi, P., & Aggarwal, A. (2006). NF-kB transcription factor: A key player in the generation of immune response. In Current Science (Vol. 90, Issue 4, pp. 519–531). www.ncbi.nlm.nih.gov/entrez/query.fcgi
Xu, X., Qiu, C., Zhu, L., Huang, J., Li, L., Fu, W., Zhang, L., Wei, J., Wang, Y., Geng, Y., Zhang, X., Qiao, W., & Xu, J. (2014). IFN-Stimulated Gene LY6E in Monocytes Regulates the CD14/TLR4 Pathway but Inadequately Restrains the Hyperactivation of Monocytes during Chronic HIV-1 Infection. The Journal of Immunology, 193(8), 4125–4136. https://doi.org/10.4049/jimmunol.1401249
Yeom, C. J., Zeng, L., Goto, Y., Morinibu, A., Zhu, Y., Shinomiya, K., Kobayashi, M., Itasaka, S., Yoshimura, M., Hur, C. G., Kakeya, H., Hammond, E. M., Hiraoka, M., & Harada, H. (2016). LY6E: A conductor of malignant tumor growth through modulation of the PTEN/PI3K/Akt/HIF-1 axis. Oncotarget, 7(40), 65837–65848. https://doi.org/10.18632/oncotarget.11670
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
5/25/2020
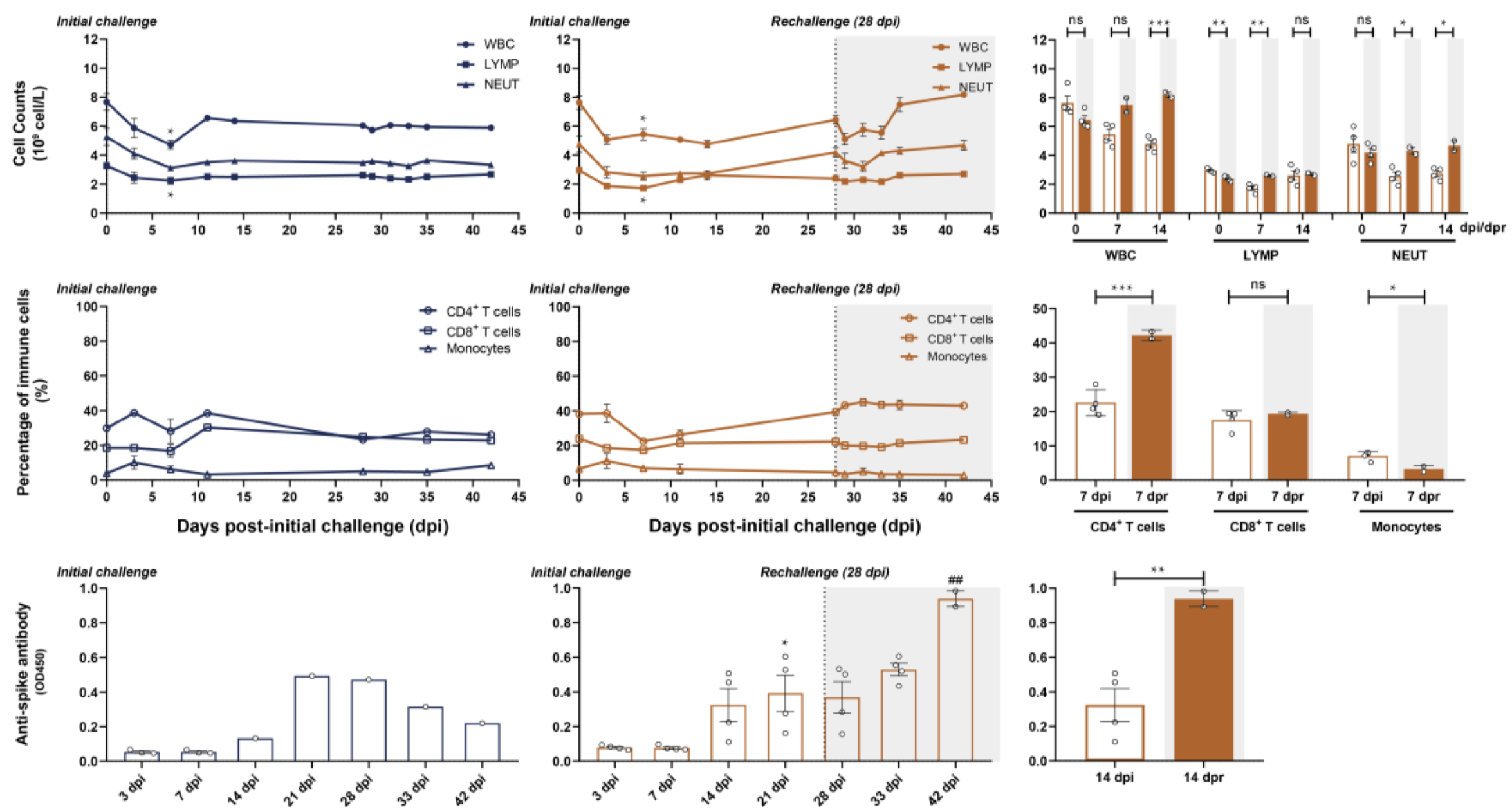
In an effort to determine the SARS-CoV-2-specific antibody response longitudinally, a team of researchers infected rhesus macaques with the virus and measured the viral load and antibody titer over the course of several weeks. Initially, six macaques were administered intratracheal injections of SARS-CoV-2 viral particles. At 28 days post infection (dpi, equal to 0 days post re-challenge [dpr]), four of the six were re-challenged intratracheally with the same viral dose; various cell and antibody counts were carried out to 45 dpi (17 dpr, Figure 1). In the primary challenge phase, peak viral load was detected by nasal and pharyngeal swabs at 3 dpi, and subsequently declined to undetectable levels at 14 dpi. Of the two macaques that were not given a secondary viral challenge, SARS-specific antibodies peaked at 21 dpi at a level that was significantly higher than the titer at 3 dpi. White blood cell (WBC), total lymphocyte, and neutrophil counts were elevated in the secondary challenged monkeys compared to the primary challenged monkeys; this difference reached significance by 14 dpi. Additionally, CD4+ T cells were significantly elevated in secondary challenged monkeys at 7 dpr compared to primary challenged monkeys at 7 dpi (Bao et al., 2020). Overall, these data demonstrate a potential build-up of specific neutralizing antibodies and immune cell response within a few weeks of viral infection that may serve ultimately to prevent reinfection by the same pathogen. There are several limitations to note: first, the antibody titers measured were not assessed for neutralizing capacity, so the anti-spike protein antibody titers may not necessarily accurately represent an infection-inhibiting immune response. Second, because neutrophils and monocytes are not antigen-specific, their presence is not necessarily indicative of a SARS-CoV-2-specific immune response. Similarly, enhanced antibody production may simply be the product of a cumulative immune response, rather than a signature of re-infection. To determine the precise mechanism, it will be useful in future studies to expand the course of the study, such that re-infection occurs far enough post-primary infection that the secondary immune response can be qualitatively and quantitatively differentiated from the initial response.
The hormones epinephrine (adrenaline), norepinephrine (noradrenaline), and dopamine comprise a class of molecules known as catecholamines. The catecholamines occupy various niches in the central and peripheral nervous systems where they act to modify behavior and as signaling molecules between neurons (Axelrod, Weinshilboum, 1972; Lotharius, Brundin, 2002; Hirasawa et al., 1993). Cytokine release syndrome (CRS) is a condition characterized by the hypersecretion of pro-inflammatory cytokines in response to antigenic stimulation and is often accompanied by a surge in catecholamines, some of which have been shown to be secreted directly by immune cells (Flierl et al., 2007). These catecholamines act in a feed-forward loop to augment the production of pro-inflammatory cytokines, which occurs primarily via the α1-type adrenergic receptor (α1-AR).
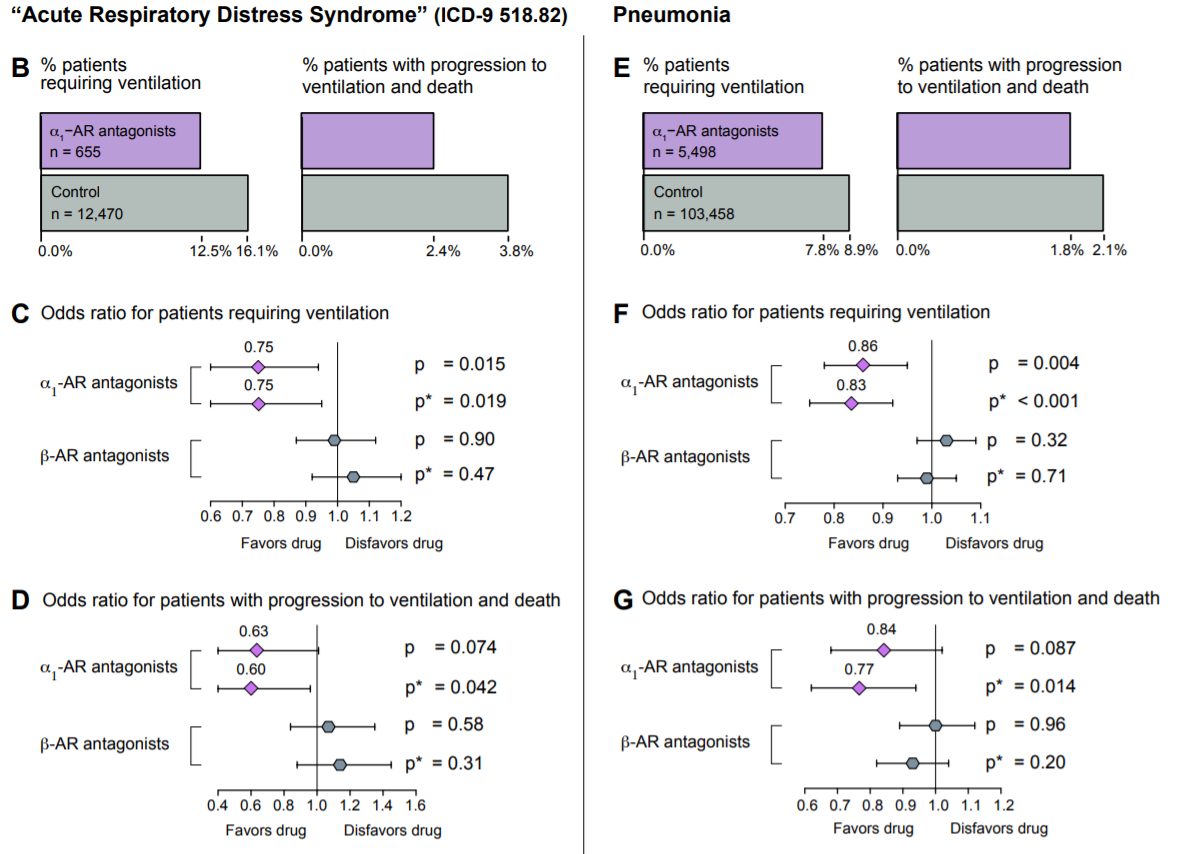
In a paper published earlier this month, Konig et al. analyzed the efficacy of α1-AR antagonists as protective agents in acute respiratory distress syndrome (ARDS) and pneumonia, two conditions that are almost ubiquitously observed in COVID-19. In this retrospective study, 122,081 patients from the MarketScan Research Database were split into an ARDS cohort and a pneumonia cohort, and were screened for α1-AR antagonist use and analyzed for one of two outcomes: progression to ventilation, or further progression to death while ventilated. Due to the retrospective nature of the study, α1-AR antagonist use was defined as “patients having filled [an] α1-AR antagonist prescription…in the year preceding [diagnosis of ARDS or pneumonia] for more than an aggregate of 180 days” (these patients would have been prescribed an antagonist for the treatment of various other chronic conditions that are not related to pneumonia or ARDS, conditions such as benign prostatic hyperplasia or hypertension). Of the 13,125 patients in the ARDS cohort, α1-AR antagonist users had a roughly 22% lower incidence of invasive mechanical ventilation compared to non-users and a roughly 36% lower incidence of further progression to death in the hospital while ventilated. In contrast, prior use of β-AR antagonists was not correlated with either outcome. Similarly, of the 108,956 patients in the pneumonia cohort, α1-AR antagonist users exhibited a roughly 13% lower incidence of invasive mechanical ventilation compared to non-users, as well as an approximately 16% lower incidence of further progression to death in the hospital on ventilation. Again, prior use of β-AR antagonists was not correlated with either outcome (Konig et al., 2020). It is important to note that though this study did not examine COVID-19 patients, it has vast implications for the use of catecholamine-interfering treatments in COVID-19, and suggests a possible path for the amelioration of the hyperinflammatory symptoms observed in these patients.
References
Axelrod, J., & Weinshilboum, R. (1972). Catecholamines. In New England Journal of Medicine (Vol. 287, Issue 5, pp. 237–242). Massachusetts Medical Society. https://doi.org/10.1056/NEJM197208032870508
Bao, L., Deng, W., Gao, H., Xiao, C., Liu, J., Xue, J., Lv, Q., Liu, J., Yu, P., Xu, Y., Qi, F., Qu, Y., Li, F., Xiang, Z., Yu, H., Gong, S., Liu, M., Wang, G., Wang, S., … Qin, C. (2020). Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. BioRxiv, 2020.03.13.990226. https://doi.org/10.1101/2020.03.13.990226
Flierl, M. A., Rittirsch, D., Nadeau, B. A., Chen, A. J., Sarma, J. V., Zetoune, F. S., McGuire, S. R., List, R. P., Day, D. E., Hoesel, L. M., Gao, H., Van Rooijen, N., Huber-Lang, M. S., Neubig, R. R., & Ward, P. A. (2007). Phagocyte-derived catecholamines enhance acute inflammatory injury. Nature, 449(7163), 721–725. https://doi.org/10.1038/nature06185
Hírasawa, A., Horie, K., Tanaka, T., Takagaki, K., Murai, M., Yano, J., & Tsujimoto, G. (1993). Cloning, Functional Expression and Tissue Distribution of Human cDNA for the α1C-Adrenergic Receptor. Biochemical and Biophysical Research Communications, 195(2), 902–909. https://doi.org/10.1006/bbrc.1993.2130
Konig, M. F., Powell, M., Staedtke, V., Bai, R.-Y., Thomas, D. L., Fischer, N., Huq, S., Khalafallah, A. M., Koenecke, A., Papadopoulos, N., Kinzler, K. W., Vogelstein, B., Vogelstein, J. T., Athey, S., Zhou, S., & Bettegowda, C. (2020). Targeting the catecholamine-cytokine axis to prevent SARS-CoV-2 cytokine storm syndrome. MedRxiv, 2, 2020.04.02.20051565. https://doi.org/10.1101/2020.04.02.20051565
Lotharius, J., & Brundin, P. (2002). Pathogenesis of parkinson’s disease: Dopamine, vesicles and α-synuclein. Nature Reviews Neuroscience, 3(12), 932–942. https://doi.org/10.1038/nrn983
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
5/18/2020
ACE2 is well established as the primary target for viral attachment and entry for both SARS-CoV-1 and SARS-CoV-2 (Li et al., 2005; Zhang et al., 2020). In addition, several other molecules have been discovered to play an accessory role in permitting viral entry into host cells. One of these accessory proteins is a molecule called TMPRSS2, a serine protease that acts to prime the viral spike (S) protein of SARS-CoV-1 prior to interaction with ACE2 and was also found to be implicated in the attachment of SARS-CoV-2 (Hoffman et al., 2020). More recently, CD147 was found to be a novel mediator of viral attachment in SARS-CoV-2 infection. CD147 is a membrane-spanning glycoprotein that is prominently associated with tumor development, protection from plasmodium invasion, and viral infection: it has been shown to inhibit signaling pathways of the tumor suppressor p53, block red blood cell infection of the protozoan that causes malaria P. falciparum, and facilitate HIV invasion (Huang et al., 2014; Zhang et al., 2017; Pushkarsky et al., 2001). Previously, Wang et al. showed that CD147 facilitates SARS-CoV-1 entry, which could be inhibited by CD147-antagonist peptide-9 in the HEK293T cell line (Chen et al., 2005). Here, they first demonstrated that humanized anti-CD147 monoclonal antibody Meplazumab competitively inhibited binding of the S protein and CD147 – compared to controls, CD147-blocked cells displayed significantly less infection by SARS-CoV-2. Additionally, Surface Plasmon Resonance (SPR, Top figure 1A) and co-immunoprecipitation with anti-CD147 and anti-S protein (Top figure 1B) confirmed the interaction of the two molecules with micromolar affinity (Kd = 1.85 x 10-7 M); and competitive direct ELISA (Top figure 1C-D) confirmed a dose-dependent interaction and its inhibition by Meplazumab. Follow-up observation by electron microscopy in infected Vero E6 cells (Bottom figure 1A-C) demonstrated significant co-localization of CD147 and S protein in viral inclusion bodies (Wang et al., 2020). The discovery of novel routes of entry provides unique opportunities for supplementing the molecular arsenal against SARS-CoV-2.

Convalescent plasma screening for IgG and IgM against pathogens is an important method for [relatively] quick identification of potentially neutralizing epitopes on those pathogens. However, of growing import is the transmission of the potentially protective IgA from mothers previously infected with SARS-CoV-2 to their babies through breastfeeding. In a recent study, fifteen milk samples were obtained from previously infected donors and compared to post-outbreak controls (the post-infection samples were obtained between 14 and 30 days after symptoms abated in these patients). In twelve of the fifteen samples, purified IgA demonstrated significant reactivity to the receptor-binding domain (RBD) of the S protein of SARS-CoV-2. Overall, the optical density measurements for the COVID-19-recovered patients were significantly greater than control measurements for IgA, IgG and secretory-type antibodies specific for the RBD (Fox et al., 2020). Growing support for the intestine’s role in proper immune system development and functioning has lent credence to a connection between this environment and that of the IgA-protected mucosa of the mammary glands – the so-called “entero-mammary link”. The intestines, being a mucosal environment, are predominant sites of IgA production and secretion by intraepithelial B cells proliferating in mesenteric lymph nodes. The authors postulate that IgA specific for SARS-CoV-2 may have originated in the gut, as previous mammary IgA has been found to exhibit specificity for enteric antigens (Seppo et al., 2017). This work suggests an important role for IgA in the immune response against SARS-CoV-2.

References
Chen, Z., Mi, L., Xu, J., Yu, J., Wang, X., Jiang, J., Xing, J., Shang, P., Qian, A., Li, Y., Shaw, P. X., Wang, J., Duan, S., Ding, J., Fan, C., Zhang, Y., Yang, Y., Yu, X., Feng, Q., … Zhu, P. (2005). Function of HAb18G/CD147 in Invasion of Host Cells by Severe Acute Respiratory Syndrome Coronavirus. The Journal of Infectious Diseases, 191(5), 755–760. https://doi.org/10.1086/427811
Fox, A., Marino, J., Amanat, F., Krammer, F., Hahn-Holbrook, J., Zolla-Pazner, S., & Powell, R. L. (n.d.). Evidence of a significant secretory-IgA-dominant SARS-CoV-2 immune response in human milk following recovery from COVID-19. https://doi.org/10.1101/2020.05.04.20089995
Hoffmann, M., Kleine-Weber, H., Schroeder, S., Krüger, N., Herrler, T., Erichsen, S., Schiergens, T. S., Herrler, G., Wu, N. H., Nitsche, A., Müller, M. A., Drosten, C., & Pöhlmann, S. (2020). SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell, 181(2), 271-280.e8. https://doi.org/10.1016/j.cell.2020.02.052
Huang, Q., Li, J., Xing, J., Li, W., Li, H., Ke, X., Zhang, J., Ren, T., Shang, Y., Yang, H., Jiang, J., & Chen, Z. (2014). CD147 promotes reprogramming of glucose metabolism and cell proliferation in HCC cells by inhibiting the p53-dependent signaling pathway. Journal of Hepatology, 61(4), 859–866. https://doi.org/10.1016/j.jhep.2014.04.035
Li, W., Zhang, C., Sui, J., Kuhn, J. H., Moore, M. J., Luo, S., Wong, S. K., Huang, I. C., Xu, K., Vasilieva, N., Murakami, A., He, Y., Marasco, W. A., Guan, Y., Choe, H., & Farzan, M. (2005). Receptor and viral deLi, W., Zhang, C., Sui, J., Kuhn, J. H., Moore, M. J., Luo, S., Wong, S. K., Huang, I. C., Xu, K., Vasilieva, N., Murakami, A., He, Y., Marasco, W. A., Guan, Y., Choe, H., & Farzan, M. (2005). Receptor and viral determinants of SARS-c. EMBO Journal, 24(8), 1634–1643. https://doi.org/10.1038/sj.emboj.7600640
Pushkarsky, T., Zybarth, G., Dubrovsky, L., Yurchenko, V., Tang, H., Guo, H., Toole, B., Sherry, B., & Bukrinsky, M. (2001). CD147 facilitates HIV-1 infection by interacting with virus-associated cyclophilin A. Proceedings of the National Academy of Sciences of the United States of America, 98(11), 6360–6365. https://doi.org/10.1073/pnas.111583198
Seppo, A. E., Savilahti, E. M., Berin, M. C., Sampson, H. A., & Järvinen, K. M. (2017). Breast milk IgA to foods has different epitope specificity than serum IgA—Evidence for entero-mammary link for food-specific IgA? Clinical and Experimental Allergy, 47(10), 1275–1284. https://doi.org/10.1111/cea.12945
Wang, K., Chen, W., Zhou, Y.-S., Lian, J.-Q., Zhang, Z., Du, P., Gong, L., Zhang, Y., Cui, H.-Y., Geng, J.-J., Wang, B., Sun, X.-X., Wang, C.-F., Yang, X., Lin, P., Deng, Y.-Q., Wei, D., Yang, X.-M., Zhu, Y.-M., … Chen, Z.-N. (2020). SARS-CoV-2 invades host cells via a novel route: CD147-spike protein. BioRxiv, 2020.03.14.988345. https://doi.org/10.1101/2020.03.14.988345
Zhang, H., Penninger, J. M., Li, Y., Zhong, N., & Slutsky, A. S. (2020). Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Medicine, 46(4), 586–590. https://doi.org/10.1007/s00134-020-05985-9
Zhang, M. Y., Zhang, Y., Wu, X. D., Zhang, K., Lin, P., Bian, H. J., Qin, M. M., Huang, W., Wei, D., Zhang, Z., Wu, J., Chen, R., Feng, F., Wang, B., Nan, G., Zhu, P., & Chen, Z. N. (2018). Disrupting CD147-RAP2 interaction abrogates erythrocyte invasion by Plasmodium falciparum. Blood, 131(10), 1111–1121. https://doi.org/10.1182/blood-2017-08-802918
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
5/11/2020
The complement system is an interdigitated array of three pathways that act as part of the innate immune response to protect against extracellular pathogens. Dysregulation or hyperactivation of the complement system is associated with various inflammatory and autoimmune conditions (Karpman et al., 2015; Ballanti et al., 2013; Jensen, 1967). The classical complement pathway consists of the hexameric protein C1q which, in combination with the serine proteases C1r and C1s, binds to antibody (IgM or IgG) attached to the surface of a pathogen. This interaction activates C1q to cleave C1r/s, initiating a cascade that results in the deposition of the protein C3b on the pathogen surface, marking it for uptake by phagocytic cells. The alternative pathway is initiated spontaneously in the blood by the constitutive cleavage of the protein C3 to C3(H2O), which leads to subsequent cleavage events resulting in the deposition of C3b on pathogen surfaces. Finally, the lectin pathway utilizes many of the same protein components as the classical pathway, but is initiated when proteins called mannose-binding lectins (MBLs) or ficolins recognize and bind carbohydrates on the surfaces of pathogens, thereby activating the serine proteases MASP-1, -2, and -3; likewise, the lectin pathway results in deposition of C3b (Noris, Remuzzi, 2013).
Gao et al. carried out a study to determine the effect that SARS-CoV-2 has on complement activation. Purified MASP-2 was tagged with a molecule called Flag that allows it to be observed in various assays. MASP-2-Flag was then incubated in the 293T cell line with MBL and mannan, the cell-surface molecule to which MBL binds for activation, with or without the SARS-CoV-2 nucleocapsid (N) protein, and the cells were assayed for MASP-2 cleavage. The researchers found that higher levels of cleaved MASP-2 fragments resulting from MASP-2 auto-activation were produced in the presence of the N protein compared to non-N protein cultures. Similarly, the binding of MASP-2 to MBL was significantly enhanced in the presence of the N protein. C4 is a protein in the complement cascade that is cleaved to form the downstream products C4a and C4b. C4 hydrolysis (cleavage with water) and C4b deposition on cell surfaces were observed to be significantly potentiated in the presence of the N protein. When the serum of COVID-19 patients was collected and tested for various complement components, C5a stuck out as being significantly increased, especially in severe patients. C5a is part of a class of molecules called anaphylatoxins – along with C3a, another product of the complement cascade – which have the ability to induce inflammation and changes in vascular permeability, allowing leukocytes to migrate to sites of infection (Hugli, Muller-Eberhard, 1978). Taken together, this research suggests the ability of SARS-CoV-2 to aggressively activate at least one pathway of the complement system via its nucleocapsid protein. This mechanism presents verification for and supplements the essentially canonical hyperinflammatory response observed in early COVID-19. Inhibitors of the complement cascade such as C1INH or monoclonal antibodies directed against components of the pathway may be instrumental in treating the hyperimmune symptoms in patients.

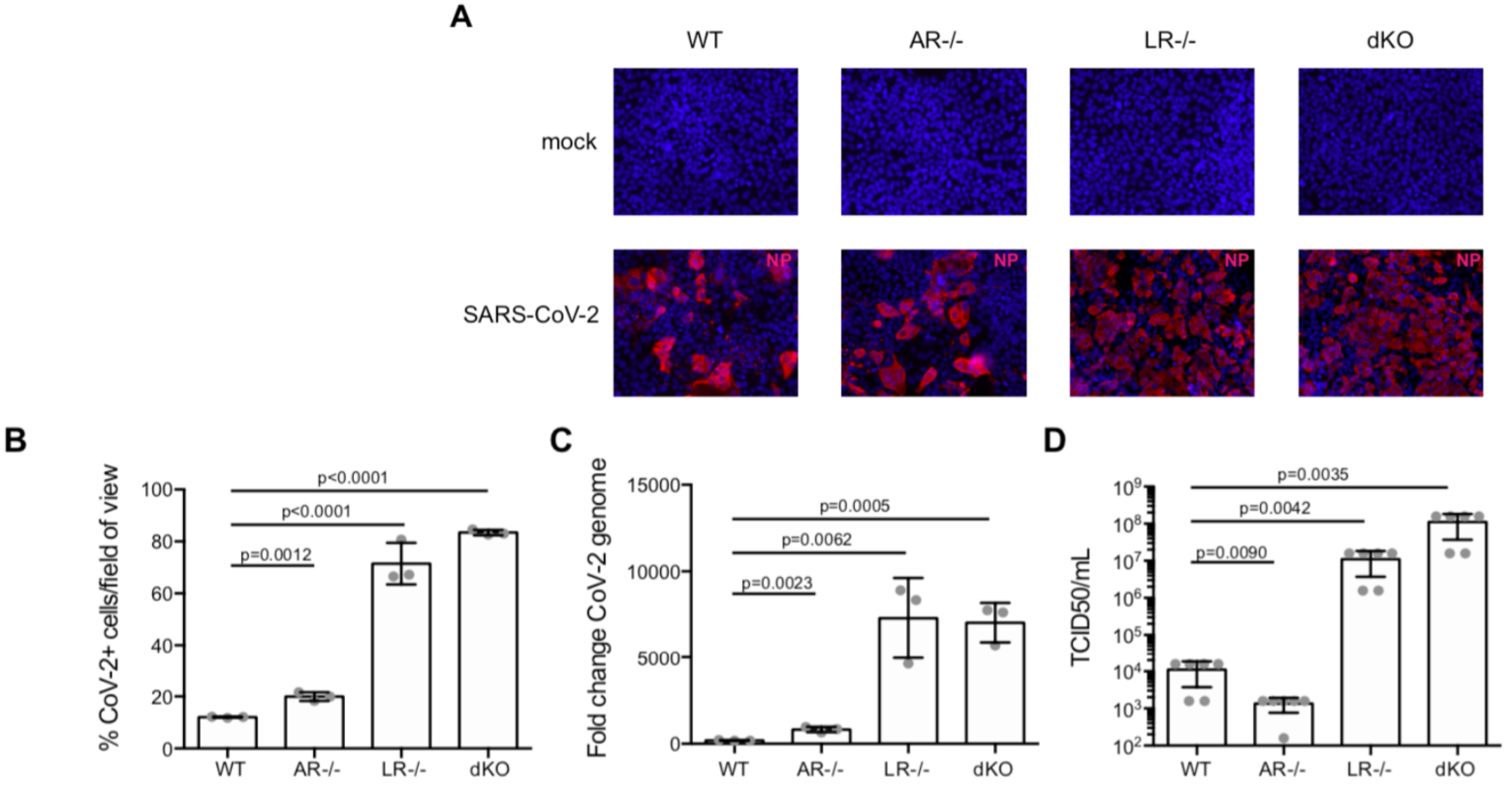
Interferon (IFN) is an important group of molecules that play a role in both innate and adaptive immunity (Ho, Armstrong, 1975). Typically, type I IFNs, secreted predominantly by plasmacytoid dendritic cells (pDCs), are thought to play the largest part in fending against and clearing viral infection. However, the roles of IFNs type II and III have also been investigated. Here, Stanifer et al. investigated the type III IFN response in intestinal epithelial cells (IECs) in response to SARS-CoV-2 infection. In this study, T84 IECs were knocked out for either the type I or type III IFN receptor, or both (double knockouts, “dKO”) and compared to wild type T84 cells that possessed functional receptors of both types. By knocking out the receptors, the researchers removed the capacity of the T84 cells to respond to type I or type III IFN. The cells were immunostained 24 hours post-infection with SARS-CoV-2 using an anti-nucleocapsid antibody. The results showed that the depletion of the type III IFN receptor resulted in a massive increase in cell infectivity, enhanced by a factor of roughly 7x compared to the type I knockouts and wild type cells. The type I knockouts exhibited only a “modest increase” in cell infectivity compared to wild type cells. In the dKO cells, an increase of roughly 7x was observed, reflecting the dysregulation of the type III IFN response in these cells. The increase in the number of infected cells upon depletion of the type III receptor was associated with significant increases in viral genome copy numbers in these cells, as well as with a 1000-fold increase in de novo synthesis of infectious viral particles. The importance of the type III IFN response in mediating infection in the intestine is thus highlighted by the significant increases in viral replication in the absence of these molecules. Future work will need to focus on characterizing the type III IFN response to SARS-CoV-2 infection in tissues outside the intestine.
A paper from just this week (5/5) details the analysis of a neutralizing antibody that could potentially lead to breakthroughs in treatments for COVID-19. A group from the Netherlands screened 51 monoclonal antibodies (mAbs) originally used against SARS-CoV-1 for cross-reactivity with SARS-CoV-2 spike protein subunits. 4 out of the 51 displayed some cross-reactivity, while only one – an mAb called 47D11 – exhibited cross-neutralizing activity. The ability of the antibody to neutralize the spike protein is one of the most important factors in determining its efficacy for therapeutic use – some antibodies may be cross-reactive and bind to the virus, but to some surface that does not prevent the viral particle from gaining entry to host cells and replicating successfully. 47D11 inhibited infection of VeroE6 cells by SARS-CoV-1 and SARS-CoV-2 in both pseudotyped (with VSV) and authentic infections. Using ELISA, 47D11 was shown to target and bind the S1B subunit of the spike protein receptor-binding domain (RBD) of both SARS-CoV-1 and SARS-CoV-2 with similar affinities (EC50 of 0.02 and 0.03 micrograms/milliliter, respectively). Of note, 47D11 binding did not compete with S1B binding to the ACE2 receptor, the known target for SARS-CoV-2 entry. Further analysis revealed that 47D11 seems to impair syncytia formation in both SARS-CoV-1 and SARS-CoV-2 infection. The mAb thus neutralizes coronavirus infections via a yet unknown mechanism. However, the cross-reactivity of 47D11 indicates that it likely targets a conserved core structure of the S1B subunit, given the fact that the binding subdomain of the spike protein is largely heterogeneous across members of the coronavirus family. This would explain the inability of the mAb to compromise the spike protein-ACE2 receptor interaction. Importantly, the existence of this mAb opens possibilities for combination monoclonal antibody treatment utilizing antibodies against various subunits of the spike protein.

References
Ballanti, E., Perricone, C., Greco, E., Ballanti, M., Di Muzio, G., Chimenti, M. S., & Perricone, R. (2013). Complement and autoimmunity. Immunologic Research, 56(2–3), 477–491. https://doi.org/10.1007/s12026-013-8422-y
Gao, T., Hu, M., Zhang, X., Li, H., Zhu, L., Liu, H., Dong, Q., Zhang, Z., Wang, Z., Hu, Y., Fu, Y., Jin, Y., Li, K., Zhao, S., Xiao, Y., Luo, S., Li, L., Zhao, L., Liu, J., … Cao, C. (2020). Highly pathogenic coronavirus N protein aggravates lung injury by MASP-2-mediated complement over-activation. MedRxiv, 2020.03.29.20041962. https://doi.org/10.1101/2020.03.29.20041962
Ho, M., & Armstrong, J. A. (1918). Interferon. 5–9. www.annualreviews.org
Hugli, T. E., & Müller-Eberhard, H. J. (1978). Anaphylatoxins: C3a and C5a. Advances in Immunology, 26(C), 1–53. https://doi.org/10.1016/S0065-2776(08)60228-X
Jensen, J. (1967). Anaphylatoxin in its relation to the complement system. Science, 155(3766), 1122–1123. https://doi.org/10.1126/science.155.3766.1122
Karpman, D., Ståhl, A. L., Arvidsson, I., Johansson, K., Loos, S., Tati, R., Békássy, Z., Kristoffersson, A. C., Mossberg, M., & Kahn, R. (2015). Complement interactions with blood cells, endothelial cells and microvesicles in thrombotic and inflammatory conditions. Advances in Experimental Medicine and Biology, 865, 19–42. https://doi.org/10.1007/978-3-319-18603-0_2
Noris, M., & Remuzzi, G. (2013). Overview of complement activation and regulation. Seminars in Nephrology, 33(6), 479–492. https://doi.org/10.1016/j.semnephrol.2013.08.001
Stanifer, M. L., Kee, C., Cortese, M., Triana, S., Mukenhirn, M., Kraeusslich, H.-G., Alexandrov, T., Bartenschlager, R., & Boulant, S. (2020). Critical role of type III interferon in controlling SARS-CoV-2 infection, replication and spread in primary human intestinal epithelial cells. BioRxiv, 2020.04.24.059667. https://doi.org/10.1101/2020.04.24.059667
Wang, C., Li, W., Drabek, D., Okba, N. M. A., Haperen, R. van, Osterhaus, A. D. M. E., Kuppeveld, F. J. M. van, Haagmans, B. L., Grosveld, F., & Bosch, B.-J. (2020). A human monoclonal antibody blocking SARS-CoV-2 infection. BioRxiv, 11(1), 2020.03.11.987958. https://doi.org/10.1101/2020.03.11.987958
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
5/4/2020
Increasingly, studies are being published that provide evidence for the tropism of SARS-CoV-2 to tissues outside the respiratory system – i.e., the affinity of SARS-CoV-2 to infect organs besides the lungs (Chu et al., 2020; Xiao, Tang et al., 2020). Of note, a postmortem study of six patients who died of COVID-19 revealed that the novel coronavirus has the potential to infect the secondary lymphoid organs – the spleen and lymph nodes (Chen, Feng et al., 2020). The secondary lymphoid organs are vital to the maintenance of the immune system and initiation of adaptive immune responses: once B and T cells mature and leave the bone marrow and thymus, respectively, they enter the follicular space of the lymph nodes and white pulp of the spleen. Here, they encounter antigens from the lymph and blood and undergo activation to become effector immune cells. This study demonstrated the expression of ACE2 – the putative target of SARS-CoV-2 – in CD68+ lymph node-resident macrophages and CD169+ splenic marginal zone-resident macrophages. Additionally, nucleocapsid protein (NP) of SARS-CoV-2 was found in splenic and lymph node tissue of all autopsied samples and was absent from healthy control splenic and lymph node tissue. Chen et al. reported that NP antigen was found only in the CD68+ and CD169+ macrophages of the autopsied samples, not in T or B cells. In the future, it will be important to determine whether NP found in these macrophages is due to infection of the spleen and lymph nodes or if it is instead the product of infection elsewhere and subsequent trafficking to the secondary lymphoid organs.
Gross examination of lymph nodes and the spleen revealed widespread necrosis and apoptosis of lymphocytes, especially in the paracortical and follicular areas of the lymph nodes. This was accompanied by a significant reduction in lymphocytes in the blood. The spleens and lymph nodes were congested and hemorrhagic across all autopsy specimens. When tested against age-matched, normal healthy controls, virally infected spleen and lymph nodes showed exaggerated lymphocyte apoptosis in autopsy specimens. This destruction of the secondary lymphoid tissues means the reduction – and possibly elimination – of the body’s ability to initiate and coordinate an adaptive immune response, leaving one lacking in the ability to fend against overwhelming infection or form pathological memory. The authors hypothesize that a persistence of viral antigen constitutively activates T and B cells, leading to a condition known as activation-induced cell death (AICD), an almost exclusively T cell phenomenon which is known to be mediated by Fas/FasL signaling (Alderson, Tough et al., 1995; Green, Droin, Pinkoski, 2003). To this end, H&E stains demonstrated significant upregulation in the expression of Fas in virally infected spleens and lymph nodes of autopsied specimens compared to controls.

A study by Boudewijns et al. characterized the effects of STAT-2-mediated secretion of type I interferons (IFNs) in Syrian hamsters infected with the human strain of SARS-CoV-2 (Boudewijns et al., 2020). STAT-2 knockout hamsters were compared against wild-type (WT) and interleukin-28 receptor (IL28R) knockout hamsters. The researchers found that the STAT-2 knockouts had higher titers of infectious virus in the lungs; relatively increased viremia; and higher levels of viral RNA in the spleen, liver, and upper and lower gastrointestinal tract. The data presented suggest that STAT-2 signaling is critical for restricting the spread of SARS-CoV-2 systemically and suppressing viral replication outside the lung compartment. Inversely, lung pathology was attenuated in STAT-2 knockouts, with limited infiltration of polymorphonuclear cells (PMNs) which correlated with the detection of only a few apoptotic bodies in the bronchial walls. WT and IL28R knockouts displayed clear signs of bronchopneumonia and peribronchial inflammation, attributed by the researchers to the infiltration of inflammatory leukocytes. This work highlights the important dual role that STAT-2 and the type I IFN response play in viral infection, the “double-edged sword” as the authors put it. Type I IFN is instrumental in fending against viral and intracellular bacterial infections by activating infected host cells to enable them to better destroy the intracellular pathogen (Durbin et al., 2000; Perry et al., 2005). Therefore, the lack of a type I IFN response mediated by knocking out the STAT-2 gene would lead to a more severe infection but attenuated inflammatory response and physiological damage, as the authors demonstrate.

References
Alderson, M. R., Tough, T. W., Davis-Smith, T., Braddy, S., Falk, B., Schooley, K. A., Goodwin, R. G., Smith, C. A., Ramsdell, F., & Lynch, D. H. (1995). Fas ligand mediates activation-induced cell death in human t lymphocytes. Journal of Experimental Medicine, 181(1), 71–77. https://doi.org/10.1084/jem.181.1.71
Boudewijns, R., Thibaut, H. J., Kaptein, S. J. F., Li, R., Vergote, V., Seldeslachts, L., Keyzer, C. De, Sharma, S., Jansen, S., Weyenbergh, J. Van, Ma, J., Martens, E., Bervoets, L., Buyten, T. Van, Jacobs, S., Liu, Y., Martí-Carreras, J., Vanmechelen, B., Wawina-Bokalanga, T., … Dallmeier, K. (2020). STAT2 signaling as double-edged sword restricting viral dissemination but driving severe pneumonia in SARS-CoV-2 infected hamsters. BioRxiv, 2020.04.23.056838. https://doi.org/10.1101/2020.04.23.056838
Chen, Y., Feng, Z., Diao, B., Wang, R., Wang, G., Wang, C., Tan, Y., Liu, L., Wang, C., Liu, Y., Liu, Y., Yuan, Z., Ren, L., & Wu, Y. (2020). The Novel Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Directly Decimates Human Spleens and Lymph Nodes. MedRxiv, 2, 2020.03.27.20045427. https://doi.org/10.1101/2020.03.27.20045427
Chu, H., Chan, J. F.-W., Yuen, T. T.-T., Shuai, H., Yuan, S., Wang, Y., Hu, B., Yip, C. C., Tsang, J. O.-L., Huang, X., Chai, Y., Yang, D., Hou, Y., Chik, K. K.-H., Zhang, X., Fung, A. Y.-F., Tsoi, H.-W., Cai, J., Chan, W.-M., … Yuen, K. (2020). Comparative tropism, replication kinetics, and cell damage profiling of SARS-CoV-2 and SARS-CoV with implications for clinical manifestations, transmissibility, and laboratory studies of COVID-19: an observational study. The Lancet Microbe, 5247(20). https://doi.org/10.1016/S2666-5247(20)30004-5
Durbin, J. E., Fernandez-Sesma, A., Lee, C.-K., Rao, T. D., Frey, A. B., Moran, T. M., Vukmanovic, S., García-Sastre, A., & Levy, D. E. (2000). Type I IFN Modulates Innate and Specific Antiviral Immunity. The Journal of Immunology, 164(8), 4220–4228. https://doi.org/10.4049/jimmunol.164.8.4220
Green, D. R., Droin, N., & Pinkoski, M. (2003). Activation-induced cell death in T cells. In Immunological Reviews (Vol. 193, Issue 1, pp. 70–81). John Wiley & Sons, Ltd. https://doi.org/10.1034/j.1600-065X.2003.00051.x
Perry, A. K., Chen, G., Zheng, D., Tang, H., & Cheng, G. (2005). The host type I interferon response to viral and bacterial infections. In Cell Research (Vol. 15, Issue 6, pp. 407–422). Nature Publishing Group. https://doi.org/10.1038/sj.cr.7290309
Xiao, F., Tang, M., Zheng, X., Liu, Y., Li, X., & Shan, H. (2020). Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. https://doi.org/10.1053/j.gastro.2020.02.055
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
4/27/2020
I reported previously on observed functional exhaustion of T cells in patients with severe COVID-19 (see 4/13 updates). In that study, Diao et al. noted increased expression of the exhaustion markers PD-1 and Tim-3 on T cells that increased with the severity of the disease and recovered to normal levels in the convalescent period (Diao et al., 2020). A similar study out of China observed significantly increased expression of another functional exhaustion marker NKG2A on CD8+ T cells as well as on natural killer (NK) cells in severe compared to non-severe cases (Zheng et al., 2020). The measurement of NKG2A was confirmed by findings of decreased intracellular CD107a, IFN-γ, IL-2, and TNF-α in both NK and CD8+ T cells. Additionally, treatment with antiviral therapy (Kaletra™) and interferon, resulted in decreased counts of NKG2A+ NK and cytotoxic T lymphocytes (CTLs) in the convalescent period in all patients observed. The functional exhaustion of immune cells – lymphocytes in particular – indicates a suppression of immune function that may play a large role in the severity of the coronavirus disease. Whether the SARS-CoV-2 virus itself mediates this suppression directly, or whether it is the result of the host inflammatory response remains to be determined. Exhaustion of NK cells in addition to T cells indicates that the immune suppression seen in COVID-19 may be more extensive than is currently understood.

As we look toward the future of COVID-19 prevention, we must take into account the nature of comorbidities and underlying genetic conditions that predispose a certain population to contracting the disease. Human Leukocyte Antigen (HLA) proteins are molecules that play the role of the major histocompatibility complex (MHC) in humans, and function in presenting antigen to the immune system for induction of an immune response (Silvestre, et al., 1970). The HLA gene family is a major contributor to what makes each person unique on a molecular/genetic level, as the polygeny and polymorphism of the class of molecules allows for expression of a unique subset in each person. Because of their direct involvement in the induction of immune responses, specific HLA alleles have been shown to be factors contributing to both susceptibility and resistance to various diseases (Hudson, Allen, 2016).
Using high-throughput proprietary screening software HLAminer™, Warren and Birol retrospectively screened bronchoalveolar lavage fluid of five patients of Chinese ethnicity who had been directly exposed to the seafood market in Wuhan at the epicenter of the outbreak. There was no classification of the severity of disease in any of the five patient samples. They discovered that four of the five patients tested expressed the HLA allele HLA-A*24:02 (Warren, Birol, 2020). The authors note that the prevalence of this specific HLA allele in the general Chinese population is known to be roughly 17.2%, a figure that is statistically significantly different from the observed 80% occurrence in the screened patients. They hypothesize that these data indicate HLA-A*24:02 to be a factor in heightened susceptibility to infection by SARS-CoV-2. Interestingly, this same allele has been described as a risk factor for the development of type I diabetes, which has itself been reported as a risk factor for severe COVID-19 (Noble, et al., 2002; Guo, et al., 2020; Fang, Karakiulakis, Roth, 2020).

Production of monoclonal antibodies (mAbs) is common as both a laboratory technique and for therapeutic use, largely against tumors (Waldmann, 1991; Weiner, Surana, Wang, 2010). To this end, mAbs developed for use against SARS-CoV and MERS-CoV were tested against the receptor binding domain (RBD) of the surface spike protein of SARS-CoV-2 (Tian, et al., 2020). The goal of the study was to determine potential candidates already in existence for use in the development of a vaccine or antiviral treatment for SARS-CoV-2. Three SARS-CoV and one MERS-CoV mAbs were tested for their affinity to the RBD of SARS-CoV-2 spike protein. It was determined that only one, SARS-CoV mAb CR3022, bound with significant affinity to SARS-CoV-2. When CR3022 was tested concurrently with ACE2 – the putative SARS-CoV-2 target – it demonstrated no competition of binding, indicating that the mAb binds the spike protein at a distinct epitope. Additionally, the monoclonal antibody m396, one of the mAbs with the highest affinity for SARS-CoV, showed no significant binding to SARS-CoV-2, highlighting the genetic and molecular distinction between the two viruses. This was postulated to be due to a difference in residues at several positions along the length of the RBD: slight discrepancies in the positions of an arginine and a tyrosine appear to play a large role in destabilizing the interaction between the RBD and m396 in SARS-CoV-2 (note the arginine at position 395 and tyrosine at position 491 [R395, Y491, respectively] in the SARS-CoV RBD and the arginine at position 408 and tyrosine at position 505 [R408, Y505] in the SARS-CoV-2 RBD in the orange model in the figure below).

References
Hudson, L. E., & Allen, R. L. (2016). Leukocyte Ig-like receptors – A Model for MHC class i disease associations. Frontiers in Immunology, 7(JUL), 281. https://doi.org/10.3389/fimmu.2016.00281
Noble, J. A., Valdes, A. M., Bugawan, T. L., Apple, R. J., Thomson, G., & Erlich, H. A. (2002). The HLA class I A locus affects susceptibility to type 1 diabetes. Human Immunology, 63(8), 657–664. https://doi.org/10.1016/S0198-8859(02)00421-4
Tian, X., Li, C., Huang, A., Xia, S., Lu, S., Shi, Z., Lu, L., Jiang, S., Yang, Z., Wu, Y., & Ying, T. (2020). Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. In Emerging Microbes and Infections (Vol. 9, Issue 1, pp. 382–385). Taylor and Francis Ltd. https://doi.org/10.1080/22221751.2020.1729069
Waldmann, T. A. (1991). Monoclonal antibodies in diagnosis and therapy. Science, 252(5013), 1657–1662. https://doi.org/10.1126/science.2047874
Warren, R. L., & Birol, I. (2020). HLA predictions from the bronchoalveolar lavage fluid samples of five patients at the early stage of the Wuhan seafood market COVID-19 outbreak. http://arxiv.org/abs/2004.07108
Weiner, L. M., Surana, R., & Wang, S. (2010). Monoclonal antibodies: Versatile platforms for cancer immunotherapy. In Nature Reviews Immunology (Vol. 10, Issue 5, pp. 317–327). Nature Publishing Group. https://doi.org/10.1038/nri2744
Zheng, M., Gao, Y., Wang, G., Song, G., Liu, S., Sun, D., Xu, Y., & Tian, Z. (2020). Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cellular & Molecular Immunology, 1–3. https://doi.org/10.1038/s41423-020-0402-2
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
4/20/2020
Updates on the intricacies of the host immunological response to SARS-CoV-2 are published daily, giving us new insights into the clinical progression of COVID-19 and potential therapeutic routes to pursue. As we move through the worst of the pandemic, it will become increasingly important to turn our attention to untangling the interactions between the novel coronavirus’ biological response and underlying comorbidities so that we may begin to treat patients more specifically. Understanding the immense overlap in the immunopathology of various pathogenic and disease processes will be critical for determining treatment plans for patients with preexisting conditions. For example, the emerging role of Th17 cells in COVID-19 has important implications due to their stimulatory function in autoimmune conditions like multiple sclerosis (Jadidi-Niaragh, Mirshafiey, 2011) and in cardiovascular diseases, which may be playing a larger role in the morbidity and mortality of COVID-19 than previously believed (Myers et al., 2016; Shi et al., 2020). Narrowing the focus of clinical treatment in patients with other afflictions can help us stay one step ahead of the virus.
Adaptive Immunity
Patients with severe COVID-19 symptoms have been observed to have a higher number of T cells that hyper-secrete interferon gamma (IFN-γ) and granulocyte-macrophage colony-stimulating factor (GM-CSF) cytokines, endowing them with a “Th1-like” function (Harbour et al., 2015). Expression of these pathogenic cells was observed to be significantly higher in severe COVID-19 compared to both non-severe and healthy control patients (Zhou et al., 2020). Notably, the same paper also identified CD14+CD16+ monocytes highly enriched in severe patients. Of note, major GM-CSF response elements and secretors of hyper-inflammatory interleukin 6 (IL-6) such as CD14+CD16+ monocytes play a key role in hyperinflammation and many autoimmune conditions. The pulmonary manifestations in COVID-19 may be due in large part to the host immune response post viral clearance and may play a role in the morbidity and mortality of patients. Given the importance of IL-6 during peak disease, anti-IL-6 or anti-IL-6R therapies are currently in clinical trials for patients experiencing “cytokine storm”.
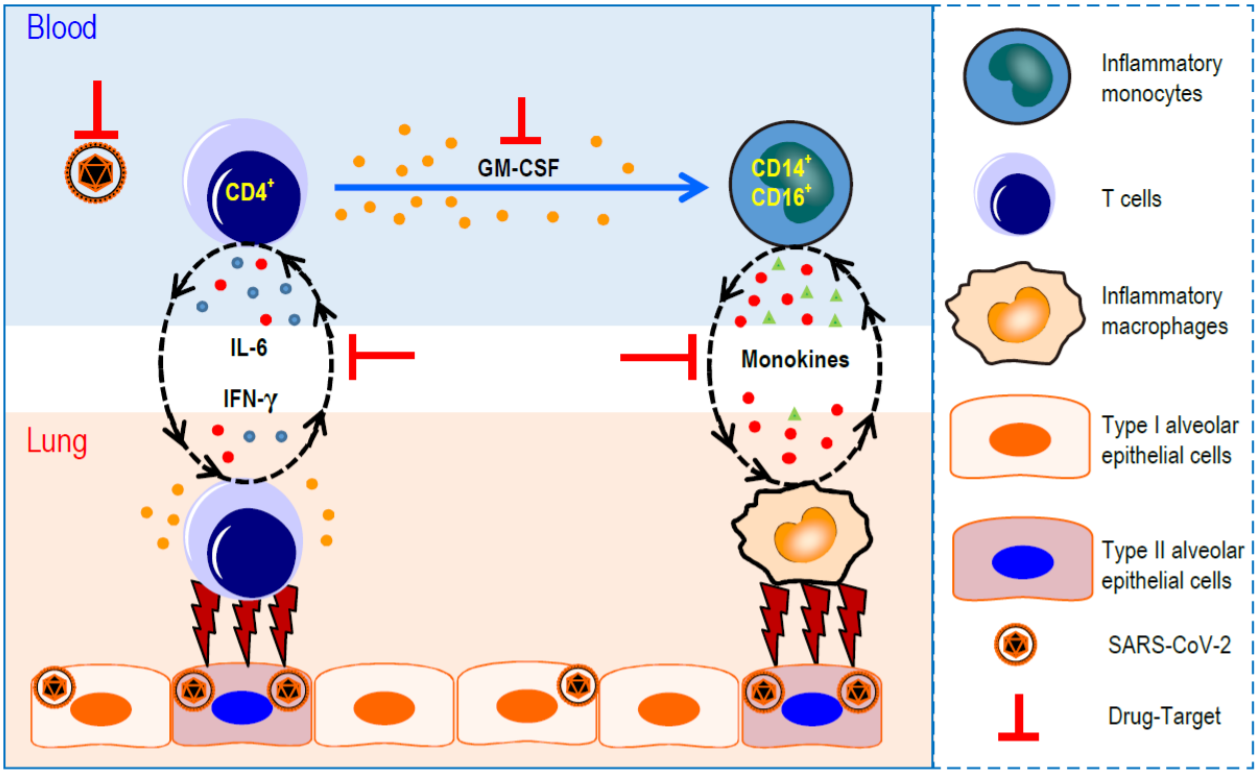
IL-6 has been implicated as a major cytokine player in the hyperinflammatory cytokine storm of SARS-CoV-2 infection (Conti et al., 2020; Qin et al., 2020; Hotez, Bottazzi, Corry, 2020). Along with IL-23, TNF-β, and a few other cytokines, IL-6 functions to stimulate the activation and differentiation of Th17 cells. The major cytokine secreted by Th17 cells, IL-17, has been shown to induce the activation of eosinophils, myeloid-derived cells involved in pro-inflammatory reactions as well as defense against parasites and some extracellular bacterial pathogens (Flier, Underhill, Weller, 1991). Several types of allergic reaction are the result of dysregulated eosinophil induction, and these allergic reactions are noted to cause pulmonary pathologies. Though the mechanism is not well understood, eosinophilia was observed in patients who received the original SARS coronavirus vaccine and was correlated with the severity of pulmonary pathology (Tseng et al., 2012). Importantly, a similar immune enhancement may result from SARS-CoV-2 immunization, resulting in hyper IL-6 production which could ultimately lead to increased Th17 responses that organize subsequent eosinophilic reactions, thereby mediating severe pulmonary immunopathology (Hotez, Bottazzi, Corry, 2020).
Innate Immunity
Pregnancy is thought to be a state of relatively immunodeficiency as well as a pro-inflammatory state under certain conditions or with certain co-morbidities. Therefore, there is a current concern about the effects of SARS-CoV-2 on pregnant patients. It is known that T and B cell counts decrease throughout pregnancy (Agaeepour, Ganio, McIlwain et al., 2017) and it is thus hypothesized that pregnant women may be more susceptible to SARS-CoV-2 infection and more prone to severe disease – i.e., the cytokine storm induced by SARS-CoV-2. Because there is no strong evidence as of yet for vertical transmission of SARS-CoV-2 from mother to fetus, the major concern remains the hyperinflammatory state of the mother which, in addition to its effects on the mother herself, has been shown to have adverse effects on fetal brain development: prolonged fever and inflammation in pregnancy have been associated with increased risks of ADHD in offspring (Werenberg Dreier et al, 2016), and excessive IL-17 was demonstrated to cause autism spectrum-like phenotypes and brain abnormalities in mice (Choi et al., 2016). The extensive overlap of immune responses and their pleiotropic effects highlight the need for timely intervention with an individualized approach.
References
Aghaeepour, N., Ganio, E. A., Mcilwain, D., Tsai, A. S., Tingle, M., Van Gassen, S., Gaudilliere, D. K., Baca, Q., McNeil, L., Okada, R., Ghaemi, M. S., Furman, D., Wong, R. J., Winn, V. D., Druzin, M. L., El-Sayed, Y. Y., Quaintance, C., Gibbs, R., Darmstadt, G. L., … Gaudilliere, B. (2017). An immune clock of human pregnancy. Science Immunology, 2(15). https://doi.org/10.1126/sciimmunol.aan2946
Choi, G. B., Yim, Y. S., Wong, H., Kim, S., Kim, H., Kim, S. V., Hoeffer, C. A., Littman, D. R., & Huh, J. R. (2016). The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science, 351(6276), 933–939. https://doi.org/10.1126/science.aad0314
Conti, P., Ronconi, G., Caraffa, A., Gallenga, C. E., Ross, R., Frydas, I., & Kritas, S. K. (2020). Induction of pro-inflammatory cytokines (IL-1 and IL-6) and lung inflammation by Coronavirus-19 (COVI-19 or SARS-CoV-2): anti-inflammatory strategies. J Biol Regul Homeost Agents, 34(2). https://doi.org/10.23812/CONTI-E
Flier, J. S., Underhill, L. H., & Weller, P. F. (1991). The immunobiology of eosinophils. In New England Journal of Medicine (Vol. 324, Issue 16, pp. 1110–1118). Massachusetts Medical Society. https://doi.org/10.1056/NEJM199104183241607
Harbour, S. N., Maynard, C. L., Zindl, C. L., Schoeb, T. R., & Weaver, C. T. (2015). Th17 cells give rise to Th1 cells that are required for the pathogenesis of colitis. Proceedings of the National Academy of Sciences of the United States of America, 112(22), 7061–7066. https://doi.org/10.1073/pnas.1415675112
Hotez, P. J., Bottazzi, M. E., & Corry, D. B. (2020). The Potential Role of Th17 Immune Responses in Coronavirus Immunopathology and Vaccine-induced Immune Enhancement. Microbes and Infection. https://doi.org/10.1016/j.micinf.2020.04.005
Jadidi-Niaragh, F., & Mirshafiey, A. (2011). Th17 Cell, the new player of neuroinflammatory process in multiple sclerosis. In Scandinavian Journal of Immunology (Vol. 74, Issue 1, pp. 1–13). John Wiley & Sons, Ltd. https://doi.org/10.1111/j.1365-3083.2011.02536.x
Liu, H., Wang, L. L., Zhao, S. J., Kwak-Kim, J., Mor, G., & Liao, A. H. (2020). Why are pregnant women susceptible to COVID-19? An immunological viewpoint. In Journal of Reproductive Immunology (Vol. 139, p. 103122). Elsevier Ireland Ltd. https://doi.org/10.1016/j.jri.2020.103122
Mor, G., Aldo, P., & Alvero, A. B. (2017). The unique immunological and microbial aspects of pregnancy. In Nature Reviews Immunology (Vol. 17, Issue 8, pp. 469–482). Nature Publishing Group. https://doi.org/10.1038/nri.2017.64
Myers, J. M., Cooper, L. T., Kem, D. C., Stavrakis, S., Kosanke, S. D., Shevach, E. M., Fairweather, D., Stoner, J. A., Cox, C. J., & Cunningham, M. W. (2016). Cardiac myosin-Th17 responses promote heart failure in human myocarditis. JCI Insight, 1(9). https://doi.org/10.1172/jci.insight.85851
Qin, C., Zhou, L., Hu, Z., Zhang, S., Yang, S., Tao, Y., Xie, C., Ma, K., Shang, K., Wang, W., & Tian, D.-S. (2020). Dysregulation of Immune Response in Patients with COVID-19 in Wuhan, China. SSRN Electronic Journal. https://doi.org/10.2139/ssrn.3541136
Sampath, P., Moideen, K., Ranganathan, U. D., & Bethunaickan, R. (2018). Monocyte Subsets: Phenotypes and Function in Tuberculosis Infection. In Frontiers in immunology (Vol. 9, p. 1726). NLM (Medline). https://doi.org/10.3389/fimmu.2018.01726
Shi, S., Qin, M., Shen, B., Cai, Y., Liu, T., Yang, F., Gong, W., Liu, X., Liang, J., Zhao, Q., Huang, H., Yang, B., & Huang, C. (2020). Association of Cardiac Injury with Mortality in Hospitalized Patients with COVID-19 in Wuhan, China. JAMA Cardiology. https://doi.org/10.1001/jamacardio.2020.0950
Tseng, C. Te, Sbrana, E., Iwata-Yoshikawa, N., Newman, P. C., Garron, T., Atmar, R. L., Peters, C. J., & Couch, R. B. (2012). Immunization with SARS coronavirus vaccines leads to pulmonary immunopathology on challenge with the SARS virus. PLoS ONE, 7(4). https://doi.org/10.1371/journal.pone.0035421
Werenberg Dreier, J., Nybo Andersen, A. M., Hvolby, A., Garne, E., Kragh Andersen, P., & Berg-Beckhoff, G. (2016). Fever and infections in pregnancy and risk of attention deficit/hyperactivity disorder in the offspring. Journal of Child Psychology and Psychiatry and Allied Disciplines, 57(4), 540–548. https://doi.org/10.1111/jcpp.12480
Zhou, Y., Fu, B., Zheng, X., Wang, D., Zhao, C., Qi, Y., Sun, R., Tian, Z., Xu, X., & Wei, H. (2020). Pathogenic T cells and inflammatory monocytes incite inflammatory storm in severe COVID-19 patients | National Science Review | Oxford Academic. National Science Review, 1–15. https://academic.oup.com/nsr/article/doi/10.1093/nsr/nwaa041/5804736
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
4/13/2020
Within the last week, more information concerning the immune mechanisms of SARS-CoV-2 has come to light. International studies characterizing the host response continue to provide updated information, and studies here in the US are beginning to gain momentum as the first wave of approval and funding reaches labs across the country. Much of what has been published concerning potential mechanisms of the viral infection have been speculations based on the previous coronaviruses (SARS-CoV-1 and MERS-CoV) and their interactions with the human immune system. With each passing week, the uniqueness of SARS-CoV-2 is being elucidated and is being used to tailor responses to the present pandemic.
Progress has been made in understanding the immunology of the infection and implementing therapies. Several follow-up convalescent plasma studies conducted in the last week have shown promise in clearing infection in patients. Details of the cellular response emerge every day and help healthcare workers and scientists redirect their efforts toward producing the most effective treatment. The next major steps include developing an understanding of how COVID-19 relates to various underlying health conditions, and how the treatment courses for those comorbidities may have to be modified to accommodate the viral infection. As we move forward, it will be more important than ever to trust the data to lead us to the best course of action.
Adaptive Immunity
Convalescent plasma trials continue to roll out of various labs around the world, and have shown great promise in treating infected patients. Despite early hesitation by the FDA and other government agencies, plasma trials are getting underway here in the US: as of April 10, 2020, the FDA approved the first convalescent plasma trial at Houston Methodist under the direction of Drs. James Musser and Eric Salazar. Yet another plasma trial out of Wuhan, China confirmed the timeframe of the IgM and IgG peaks during the host COVID-19 response (Zhou et al., 2020). The data presented by Zhou et al. indicate that the antibody (Ab) response to SARS-CoV-2 indeed follows the pattern seen in most other pathological infections, with specific, low-affinity IgM peaking within a few days of infection and high affinity IgG peaking within the first two weeks. This timeframe highlights the critical period in the first few days post-infection in which it may be necessary to treat patients more aggressively to buy time for the adaptive immune response to mount. Similarly, the first roughly 10 days post-infection offer the most efficient collection of serum from infected patients for use in transfusional therapies, as IgG seems to peak near two weeks.

As reported last week, the primary goal of many early studies was to broadly characterize the immunological response infection as a first line in directing treatment (Zhang et al., 2020; Liu et al., 2020; Bermejo-Martin et al., 2020). Increasingly, however, specifics of the response are being elucidated. The Qin lab in China noted severe lymphopenia – a reduction in the levels of T cells, B cells, and natural killer (NK) cells – as has been described previously for both severe and non-severe infections [see April 3rd updates for details], but further reported that T cells, specifically helper T (Th) cells, seem to be more drastically affected than B or NK cells (Qin et al., 2020). Moreover, it appears that the ratio of naive Th cells to memory (mature) Th cells is increased relative to controls. This could be accounted for by the fact that the virus tends to target activated Th cells, or by the fact that it prevents naive Th cells from being activated in the first place. In either case, this prevents a fully efficacious adaptive immune response from being mounted, as Th cells are the preeminent coordinators of most immune reactions.
Diao et al. published findings that three specific cytokines (interleukin-6 (IL-6), IL-10, and tumor necrosis factor alpha (TNF-α)) are not only significantly increased in infected patients compared to controls, but are also negatively correlated with total T cell, CD8+ T cell, and CD4+ T cell counts. A pseudo-longitudinal study over the course of care for their study participants noted that as T cell levels began to increase in the recovery period, levels of the cytokines mentioned above decreased. Further, they analyzed the expression of exhaustion markers on T cells, and found PD-1 and Tim-3 to be increasingly expressed as patients progressed through the disease course (Diao, et al., 2020). Functional exhaustion is the progressive loss of function that occurs as cells perform their duties and become “worn out” over time, sometimes culminating in death of the cell (Yi, Cox, Zajac, 2010). In the context of COVID-19, this presents a potential route for supplementing the immune response by treating infected patients with anti-PD-1 or anti-Tim-3 therapies. Diao et al. hypothesize that hypersecretion of IL-6, IL-10 and TNF-α is responsible for the upregulation of exhaustion markers in response to SARS-CoV-2; immunomodulators or suppressants may be key in preventing immunological exhaustion.
Innate Immunity
The focus of innate immunological studies is to understand the cytokine storm and to tease apart its specific elements. A retrospective study of 452 patients infected with SARS-CoV-2 in Shanghai, China. demonstrated that patients with severe cases of the disease present with lymphopenia, as described previously, as well as lower percentages of myeloid cells including monocytes, eosinophils and basophils when compared with non-severe cases. Additionally, several non-cytokine biomarkers were observed, including procalcitonin and serum ferritin (Qin et al., 2020). These biomarkers represent potential indicators of disease progression, similar to the cytokines reported previously (Zhang et al., 2020; Liu et al., 2020). The exact mechanism of the cytokine storm/superantigenic component of SARS-CoV-2 remains to be determined. Understanding the details of the innate response is important for planning treatment regimens for those with underlying comorbidities, as the hyperactive immune response has been implicated as a major causal factor of the pathology of the infection.
References
Abdulamir, A. S., & Hafidh, R. R. (2020). The Possible Immunological Pathways for the Variable Immunopathogenesis of COVID—19 Infections among Healthy Adults, Elderly and Children. Electronic Journal of General Medicine, 17(4), em202. https://doi.org/10.29333/ejgm/7850
Bermejo-Martin, J. F., Almansa, R., Menéndez, R., Mendez, R., Kelvin, D. J., & Torres, A. (2020). Lymphopenic community acquired pneumonia as signature of severe COVID-19 infection. Journal of Infection, 0(0). https://doi.org/10.1016/j.jinf.2020.02.029
Diao, B., Wang, C., Tan, Y., Chen, X., Liu, Y., Ning, L., Chen, L., Li, M., Liu, Y., Wang, G., Yuan, Z., Feng, Z., Wu, Y., & Chen, Y. (2020). Reduction and Functional Exhaustion of T Cells in Patients with Coronavirus Disease 2019 (COVID-19). MedRxiv, 2020.02.18.20024364. https://doi.org/10.1101/2020.02.18.20024364
Liu, H., Wang, L. L., Zhao, S. J., Kwak-Kim, J., Mor, G., & Liao, A. H. (2020). Why are pregnant women susceptible to COVID-19? An immunological viewpoint. In Journal of Reproductive Immunology (Vol. 139, p. 103122). Elsevier Ireland Ltd. https://doi.org/10.1016/j.jri.2020.103122
Qin, C., Zhou, L., Hu, Z., Zhang, S., Yang, S., Tao, Y., Xie, C., Ma, K., Shang, K., Wang, W., & Tian, D.-S. (2020). Dysregulation of Immune Response in Patients with COVID-19 in Wuhan, China. SSRN Electronic Journal. https://doi.org/10.2139/ssrn.3541136
Zhang, J. jin, Dong, X., Cao, Y. yuan, Yuan, Y. dong, Yang, Y. bin, Yan, Y. qin, Akdis, C. A., & Gao, Y. dong. (2020). Clinical characteristics of 140 patients infected with SARS-CoV-2 in Wuhan, China. Allergy: European Journal of Allergy and Clinical Immunology. https://doi.org/10.1111/all.14238
Zheng, M., Gao, Y., Wang, G., Song, G., Liu, S., Sun, D., Xu, Y., & Tian, Z. (2020). Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cellular & Molecular Immunology, 1–3. https://doi.org/10.1038/s41423-020-0402-2
Zhou, P., Yang, X. Lou, Wang, X. G., Hu, B., Zhang, L., Zhang, W., Si, H. R., Zhu, Y., Li, B., Huang, C. L., Chen, H. D., Chen, J., Luo, Y., Guo, H., Jiang, R. Di, Liu, M. Q., Chen, Y., Shen, X. R., Wang, X., … Shi, Z. L. (2020). A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature, 579(7798), 270–273. https://doi.org/10.1038/s41586-020-2012-7
COVID-19 Immunology Updates
Written by: Parker Davis
Edited by: Jina Zhou and Esther Melamed
4/7/2020
The host immune response to SARS-CoV-2 infection has been documented and characterized to some extent over the course of the last few months. Still in the early stages of understanding, there is some contention about the exact mechanism of the coronavirus infection. The earliest retrospective studies characterized a cytokine storm of pro-inflammatory mediators, suggesting a superantigenic component of the novel coronavirus. More recent papers, however, have demonstrated a more nuanced view of the disease progression, even noting a dampening effect on some subsets of T cells and other innate cells (Huang, et al., 2020). The particular mechanism by which SARS-CoV-2 mediates this dampening effect is not yet understood, though it may be the case that it utilizes a pathway similar to that of previous coronaviruses (Prompetchara, et al., 2020).
Despite the uncertainty of the host response, impressive progress todate has been made toward developing a vaccine and identifying clinical manifestations that could potentially serve as biomarkers of disease progression and severity. Several studies have looked at differences between ICU and non-ICU patient cases and scientists have begun to tease apart the immunological profile at different stages of infection (Liu, et al., 2020). Similarly, immune responses to potential therapies have been characterized, and the knowledge gleaned will be useful in directing care for infected patients, especially those with underlying conditions that may be affected by the host response to infection. Much more information will become available as more and more researchers turn their attention to stemming the spread of the COVID-19 disease.
Adaptive Immunity
Convalescent plasma trials have shown promise as potential therapy for SARS-CoV-2 infection, both in vitro and in vivo. A preliminary study out of Shenzhen, China treated 5 “critically ill” patients presenting with COVID-19 and acute respiratory distress syndrome (ARDS) with convalescent plasma (no controls), showing marked reduction in organ failure (SOFA scores), harmful proinflammatory cytokine hypersecretion, and pulmonary lesions, with increased specific antibody titers, indicating successful transfusion and the safety of plasma from previously infected patients. ARDS also resolved in 4 out of the 5 patients, likely as a result of cleared infection. Future testing against control cases will be necessary before plasma therapy can be considered a viable treatment option. (Shen, et al., 2020).

So far, five CTL and eight B cell epitopes have been identified as potential targets for vaccines. The epitopes exhibit non-covalent interactions with MHC I/II molecules, demonstrating their efficacy as presentable antigens by professional antigen-presenting cells (APCs: B cells, dendritic cells, macrophages). Artificial neural network software was used to identify and score potential epitopes within the full amino acid sequence of the surface glycoprotein of SARS-CoV-2. All the epitopes identified but one (a CTL epitope) were found to be at least partially unique to SARS-CoV-2. Of note, further characterization will be required to determine whether mapping of these epitopes is complete given the transient covalent attachment of glycans to some epitopes in question. The immediate implementation of these epitopes into vaccines is important due to the high rates of mutation seen in typical RNA viruses. (Baruah, Bose, 2020).
Th1 and Th17 cells have been implicated in immune response in various ways, though the connections are not strong yet. Importantly, chloroquine – and hydroxy-chloroquine to a lesser extent – has been shown to induce IL-23 expression in monocyte-derived cells in the skin, stimulating secretion of IL-17 subsets by CD4+ T cells. COVID-19 therapies targeting these pathways will be very important in patients with multiple sclerosis and other autoimmune conditions characterized by Th1/Th17 responses. (Prompetchara, et al., 2020).
A study (n = 16) from the Sichuan Province of China characterizing the adaptive immune response to COVID-19 has shown that several clusters of CD8+ and dendritic cells are increased, and that B cell and helper T cell clusters seem to be downregulated. Specifically, interferon-producing plasmacytoid dendritic cells are upregulated in the response, and the interferon-MAPK pathway has been suggested as one of the primary response pathways in host cells, though the details are still unclear. If this is the case, supplementation with exogenous interferon may further stimulate and enhance the immune response to SARS-CoV-2 infection. The study also performed a preliminary profile of TCR and BCR expression in vivo and demonstrated that severe cases of COVID-19 seem to be related to strong autoimmune responses, potentially indicating some cross-reactivity between human and viral antigens. (Huang, et al., 2020).
Innate Immunity
In general, infected patients present with lymphopenia and a cytokine storm of proinflammatory cytokines – IL-6, TNFa, IL-1B, C-reactive protein – suggesting a super antigen component of SARS-CoV-2. There has also been a significant difference documented between severe – admitted to the ICU – and non-severe cases in the levels of these cytokines. The cytokine response may therefore be a potential biomarker of disease progression and severity and may serve to inform treatment plans specific to a patient. (Zhang, et al., 2020). A study looked specifically at hypoalbuminemia, lymphopenia, and percentage of total neutrophil count among other indicators and found multiple specific measurements to be highly indicative of disease severity. (Liu, et al., 2020).

The disease presentation of lymphocytopenia was seen previously in the MERS outbreak of 2011 as a result of necrosis/apoptosis induced by the MERS coronavirus. It is hypothesized that SARS-CoV-2 may utilize a similar mechanism to dampen the immune response. The presentation of lymphocytopenia with hypercytokinemia is characteristic of lymphopenic community acquired pneumonia (L-CAP). The degree of L-CAP has been correlated with disease severity, increased mortality and dysregulated immune responses. Endothelial dysfunction has been hypothesized as a primary predisposing factor to low lymphocyte/high cytokine counts leading to more severe disease and novel coronavirus pneumonia (NCP). Drugs targeting endothelial dysfunction – leakage or tearing, etc. – and lymphocyte apoptosis are potential therapies for mitigating disease progression to more severe stages. (Bermejo-Martin, et al., 2020).
References
Baruah, V., & Bose, S. (2020). Immunoinformatics-aided identification of T cell and B cell epitopes in the surface glycoprotein of 2019-nCoV. Journal of Medical Virology, 92(5), 495–500. https://doi.org/10.1002/jmv.25698
Bermejo-Martin, J. F., Almansa, R., Menéndez, R., Mendez, R., Kelvin, D. J., & Torres, A. (2020). Lymphopenic community acquired pneumonia as signature of severe COVID-19 infection. Journal of Infection, 0(0). https://doi.org/10.1016/j.jinf.2020.02.029
Huang, L., Shi, Y., Gong, B., Jiang, L., Liu, X., Yang, J., Tang, J., You, C., Jiang, Q., Long, B., Zeng, T., Luo, M., Zeng, F., Zeng, F., Wang, S., Yang, X., & Yang, Z. (2020). Blood single cell immune profiling reveals the interferon-MAPK pathway mediated adaptive immune response for COVID-19. MedRxiv, 2020.03.15.20033472. https://doi.org/10.1101/2020.03.15.20033472
Liu, Y., Yang, Y., Zhang, C., Huang, F., Wang, F., Yuan, J., Wang, Z., Li, J., Li, J., Feng, C., Zhang, Z., Wang, L., Peng, L., Chen, L., Qin, Y., Zhao, D., Tan, S., Yin, L., Xu, J., … Liu, L. (2020). Clinical and biochemical indexes from 2019-nCoV infected patients linked to viral loads and lung injury. Science China Life Sciences, 63(3), 364–374. https://doi.org/10.1007/s11427-020-1643-8
Prompetchara, E., Ketloy, C., & Palaga, T. (2020). Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. Asian Pacific Journal of Allergy and Immunology. https://doi.org/10.12932/AP-200220-0772
Shen, C., Wang, Z., Zhao, F., Yang, Y., Li, J., Yuan, J., Wang, F., Li, D., Yang, M., Xing, L., Wei, J., Xiao, H., Yang, Y., Qu, J., Qing, L., Chen, L., Xu, Z., Peng, L., Li, Y., … Liu, L. (2020). Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA. https://doi.org/10.1001/jama.2020.4783
Zhang, J. Jin, Dong, X., Cao, Y. yuan, Yuan, Y. Dong, Yang, Y. bin, Yan, Y. Qin, Akdis, C. A., & Gao, Y. Dong. (2020). Clinical characteristics of 140 patients infected with SARS-CoV-2 in Wuhan, China. Allergy: European Journal of Allergy and Clinical Immunology. https://doi.org/10.1111/all.14238