Selected Publications
Osorio D, McGrail DJ, Sahni N and Yi S#. (2023) Scalable drug combination prioritization framework for precision oncology using single-cell based transfer learning. Under revision. (#corresponding author) Preprint: https://www.biorxiv.org/content/10.1101/2022.04.06.487357v1
Pan X, Yun J, Coban Akdemir ZH, Jiang X, Sahni N, Wu E, Huang JH and Yi S#. (2023) AI-DrugNet: A network-based deep learning model for drug repurposing and combination therapy in neurological disorders. Comput Struct Biotechnol J. 21, 1533-42. (#corresponding author)
Amici DR, Cingoz H, Alasady MJ, Alhayek S, Phoumyvong CM, Sahni N, Yi S and Mendillo ML. (2023) The HAPSTR2 retrogene buffers stress signaling and resilience in mammals. Nature Communications. 14, 152.
Hua X, Li Y, Pentaparthi SR, McGrail DJ, Zou R, Guo L, Shrawat A, Cirillo KM, Li Q, Bhat A, Xu M, Qi D, Singh A, McGrath F, Andrews S, Aung KL, Das J, Zhou Y, Lodi A, Mills GB, Eckhardt SG, Mendillo ML, Tiziani S, Wu E, Huang JH, Sahni N and Yi S#. (2023) Landscape of microRNA regulatory network architecture and functional rerouting in cancer. Cancer Research. 83, 59-73. (#corresponding author)
Pan X, Coban Akdemir ZH, Gao R, Jiang X, Sheynkman GM, Wu E, Huang JH, Sahni N and Yi S#. (2023) AD-Syn-Net: Systematic identification of Alzheimer’s disease-associated mutation and co-mutation vulnerabilities via deep learning. Brief Bioinformatics. 2023, 1–12. bbad030. (#corresponding author)
Chakravarty AK, McGrail DJ, Lozanoski TM, Dunn BS, Shih DJ, Cirillo KM, Cetinkaya S, Zheng WJ, Mills GB, Yi S#, Jarosz DF# and Sahni N#. (2022) Biomolecular condensation: a new phase in cancer research. Cancer Discovery. 12, 2031-43. (#co-corresponding author)
Shin W, Su Z, Yi S# and Kim HJ#. (2022) Single-cell transcriptomic mapping of intestinal epithelium that undergoes 3D morphogenesis and mechanodynamic stimulation in a gut-on-a-chip. iScience. 25, 105521. (#co-corresponding author)
Pan X, Burgman B, Wu E, Huang JH, Sahni N and Yi S#. (2022) i-Modern: Integrated multi-omics network model identifies potential therapeutic targets in glioma by deep learning with interpretability. Comput Struct Biotechnol J. 20, 3511-21. (#corresponding author)
Li Y, Burgman B, Khatri IS, Pentaparthi SR, Su Z, McGrail DJ, Li Y, Wu E, Eckhardt SG, Sahni N and Yi S#. (2021) e-MutPath: Computational modelling reveals the functional landscape of genetic mutations rewiring interactome networks. Nucleic Acids Research. 49, e2. (#corresponding author)
Guo L, Li Y, Marick RA, Su Z, Yin X, Hua X, Mills GB, Sahni N and Yi S#. (2021) mi-IsoNet: Systems-scale microRNA landscape reveals rampant isoform-mediated gain of target interaction diversity and signaling specificity. Briefings in Bioinformatics. 22, bbab091. (#corresponding author)
Nizamutdinov D, Qi X, Berman MH, Dougal G, Dayawansa S, Wu E, Yi S, Stevens AB and Huang JH. (2021) Transcranial near infrared light stimulations improve cognition in patients with dementia. Aging and Disease. 12, 954-963.
Li Y, Burgman B, McGrail DJ, Sun M, Qi D, Shukla SA, Wu E, Capasso A, Lin SY, Wu CJ, Eckhardt SG, Mills GB, Li B, Sahni N and Yi S#. (2020) Integrated genomic characterization of human immunome in cancer. Cancer Research. 80, 4854-4867. (#corresponding author)
Li Y, Zhang Y, Li X, Yi S# and Xu J#. (2019) Gain-of-function mutations: An emerging advantage for cancer biology. Trends in Biochem. Sci. 44, 659-674. (#co-corresponding author)
Li Y, McGrail DJ, Xu J, Li J, Liu NN, Sun M, Lin R, Pancsa R, Zhang J, Lee JS, Wang H, Mills GB, Li X, Yi S# and Sahni N#. (2019) MERIT: Systematic analysis and characterization of Mutational Effect on RNA Interactome Topology. Hepatology 70, 532-546. (#co-corresponding author)
Li Y, McGrail DJ, Xu J, Mills GB, Sahni N and Yi S#. (2018) Gene regulatory network perturbation by genetic and epigenetic variation. Trends in Biochemical Sciences 43, 576-592. (#corresponding author)
Ng PK, Li J, Jeong KJ, Shao S, Chen H, Tsang S, Sengupta S, Wang Z, Bhavana VH, Tran R, Soewito S, Minussi D, Moreno D, Dogruluk T, Lu H, Zeng J, Ip CK, Ju Z, Wester M, Yu S, Li Y, Vellano C, Gao J, Schultz N, Karchin R, Ding L, Lu Y, Cheung LW, Chen K, Scott KL, Yi S#, Sahni N#, Liang H# and Mills GB. (2018) Systematic functional annotation of somatic mutations in cancer. Cancer Cell 33, 450-462. (#co-corresponding author)
Li Y, Sahni N, Pancsa R, McGrail DJ, Xu J, Hua X, Coulombe-Huntington J, Ryan M, Tychhon B, Sudhakar D, Hu L, Tyers M, Jiang X, Lin SY, Babu MM, and Yi S#. (2017) Revealing the determinants of widespread alternative splicing perturbation in cancer. Cell Rep. 21, 798-812. (#corresponding author)
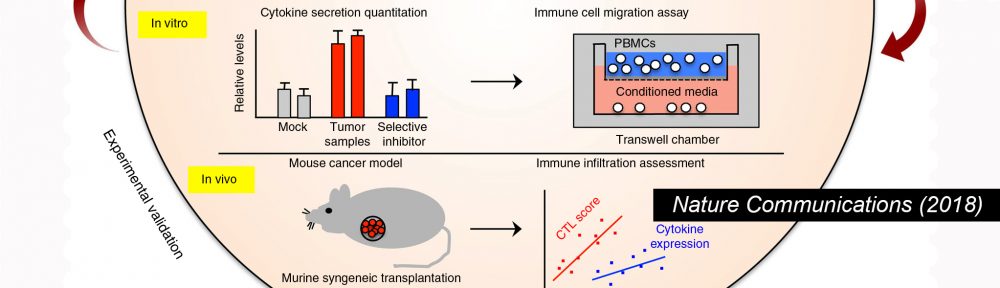
McGrail DJ, Federico L, Li Y, Dai H, Lu Y, Mills GB, Yi S#, Lin SY# and Sahni N#. (2018) Multi-omics analysis reveals neoantigen-independent immune cell infiltration in copy-number driven cancers. Nature Commun. 9, 1317. (#co-corresponding author)
Li Y, Li L, Wang Z, Pan T, Sahni N, Jin X, Wang G, Li J, Zheng X, Zhang Y, Xu J#, Yi S#, Li X#. (2018) LncMAP: Pan-cancer atlas of long noncoding RNA-mediated transcriptional network perturbations. Nucleic Acids Res. 46, 1113-23. (#co-corresponding author)
Yi S#, Lin S, Li Y, Zhao W, Mills GB and Sahni N. (2017) Functional variomics and network perturbation: Connecting genotype to phenotype in cancer. Nature Rev. Genet. 18, 395-410. (journal cover) (#corresponding author)
Yi S#, Liu NN, Hu L, Wang H and Sahni N. (2017) Base-resolution stratification of cancer mutations using functional variomics. Nature Protoc 12, 2323-41. (#corresponding author)
Karras GI, Yi S, Sahni N, Fischer M, Xie J, Vidal M, D’Andrea AD, Whitesell L and Lindquist S. (2017) HSP90 shapes the consequences of human genetic variation. Cell 168, 856-866.
Sahni N, Yi S*, Taipale M, Fuxman Bass JI, Coulombe-Huntington J, Yang F, Peng J, Weile J, Karras GI, Wang Y, et al. (2015) Widespread macromolecular interaction perturbations in human genetic disorders. Cell 161, 647-660. (*co-first author)
Barrera LA, Vedenko A, Kurland JV, Rogers JM, Gisselbrecht SS, Rossin EJ, Woodard J, Mariani L, Kock KH, Inukai S, Siggers T, Shokri L, Gordân R, Sahni N, Cotsapas C, Hao T, Yi S, Kellis M, Daly MJ, Vidal M, Hill DE, Bulyk ML. (2016) Survey of variation in human transcription factors reveals prevalent DNA binding changes. Science 351,1450-1454.
Fuxman Bass JI, Sahni N, Shrestha S, Garcia-Gonzalez A, Mori A, Bhat N, Yi S, Hill DE, Vidal M and Walhout AJ. (2015) Human gene-centered transcription factor networks for enhancers and disease variants. Cell 161, 661-673.
Rolland T, Taşan M, Charloteaux B, Pevzner SJ, Zhong Q, Sahni N, Yi S*, Lemmens I, Fontanillo C, Mosca R, et al. (2014) A proteome-scale map of the human interactome network. Cell 159, 1212-1226. (*co-first author)
Yi S, Sahni N, Daniels KJ, Lu KL, Srikantha T, Huang G, Garnaas AM and Soll DR. (2011) Alternative mating type configurations (a/alpha versus a/a or alpha/alpha) of Candida albicans result in alternative biofilms regulated by different pathways. PLoS Biol. 9, e1001117. ● Highlighted in a Research Highlight: Microbiology: Biofilm for yeast sex. Nature. 476, 129, Aug 2011
Wessels D, Srikantha T, Yi S, Kuhl S, Aravind L and Soll DR. (2006) The Shwachman-Bodian-Diamond syndrome gene encodes an RNA-binding protein that localizes to the pseudopod of Dictyostelium amoebae during chemotaxis. J. Cell Sci. 119, 370-379. ● Highlighted in a “Research Highlights” section in the journal Nature, 26 Jan 2006, 439, 372-373.