Keiler Lab
Ribosome Rescue & New Antibiotics · UT Austin
Our mission is to understand how protein quality control is maintained during stress responses and homeostasis. Most of our work addresses how stalled ribosomes are rescued by trans-translation and alternative pathways. Our goal is to characterize the fundamental biochemistry, genetics, and cell biology of these systems, and to use this knowledge to develop antibiotics and tools for basic research.
Protein Quality Control & trans-Translation
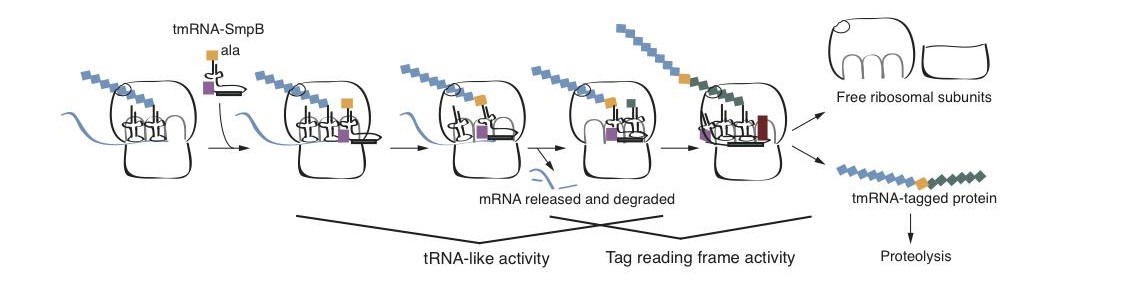
There are a wide variety of errors that can occur during transcription and translation, and cells must have mechanisms to ameliorate the deleterious effects of these errors in order to maintain optimal fitness. In bacteria, problems during transcription or translation frequently lead to a ribosome that reads to the 3′ end of an mRNA without terminating translation at a stop codon. To deal with these “non-stop” translation complexes, bacteria have a pathway known as trans-translation. During trans-translation, tmRNA, a specialized RNA molecule, and SmpB, a small protein, recognize a non-stop translation complex and use a short reading frame within tmRNA to release the ribosome and promote degradation of the nascent polypeptide. Genes encoding tmRNA and SmpB have been identified in >99% of sequenced bacterial genomes, suggesting that trans-translation confers a strong selective advantage in most environments that support bacterial life. In fact, trans-translation is essential in many bacteria. Species in which tmRNA can be deleted have a backup system that can perform similar functions, albeit less efficiently.