Written by: Hector Fernandez
Edited by: Esther Melamed
As the coronavirus pandemic continues, experts worldwide have maintained efforts toward mitigating the current situation and addressing many of the questions surrounding the outbreak of the virus. Among the pieces of the puzzle to be deciphered are aspects of the nature of the virus such as its origin, physical/biological properties, transmission, and effects on the human host. As the research efforts continue, the scientific community aims to shed light on these aspects of the virus in order to develop effective therapies, and vaccines, to treat those infected by SARS-CoV-2.
The following article gives an overview of the virology of the SARS-CoV-2 virus in hopes to inform the reader about what is currently known about the virus and other coronaviruses.
Classification
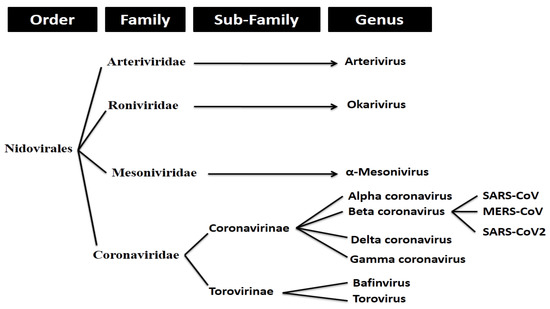
SARS-CoV-2 (more commonly referred to as ‘coronavirus’) is an enveloped, positive stranded RNA virus that forms part of a group of viruses known as beta-coronaviruses (β-CoVs). Coronaviruses make up the largest group of viruses in the Nidovirales order, which includes the families of viruses Coronaviridae, Arteriviridae, Mesoniviridae, and Roniviridae.1 Coronaviruses, referred to as Coronavirinae, are classified as a subfamily of the Coronaviridae family of viruses (with the other being the Torovirinae subfamily).
Coronavirinae viruses are further divided into four genera. These include the alpha, beta, gamma and delta coronaviruses. As previously mentioned, SARS-CoV-2 forms part of the beta coronavirus genus.
Genomic Organization
The SARS-CoV-2 genome is made up of a single-stranded positive-sense RNA (+ssRNA) molecule whose size varies from 29.8 kb to 29.9 kb (3). The genome consists of a 5’ cap domain along with a 3̘’poly (A) tail. Therefore, the viral RNA can act like an mRNA molecule ready for translation by the host cell without further modification.
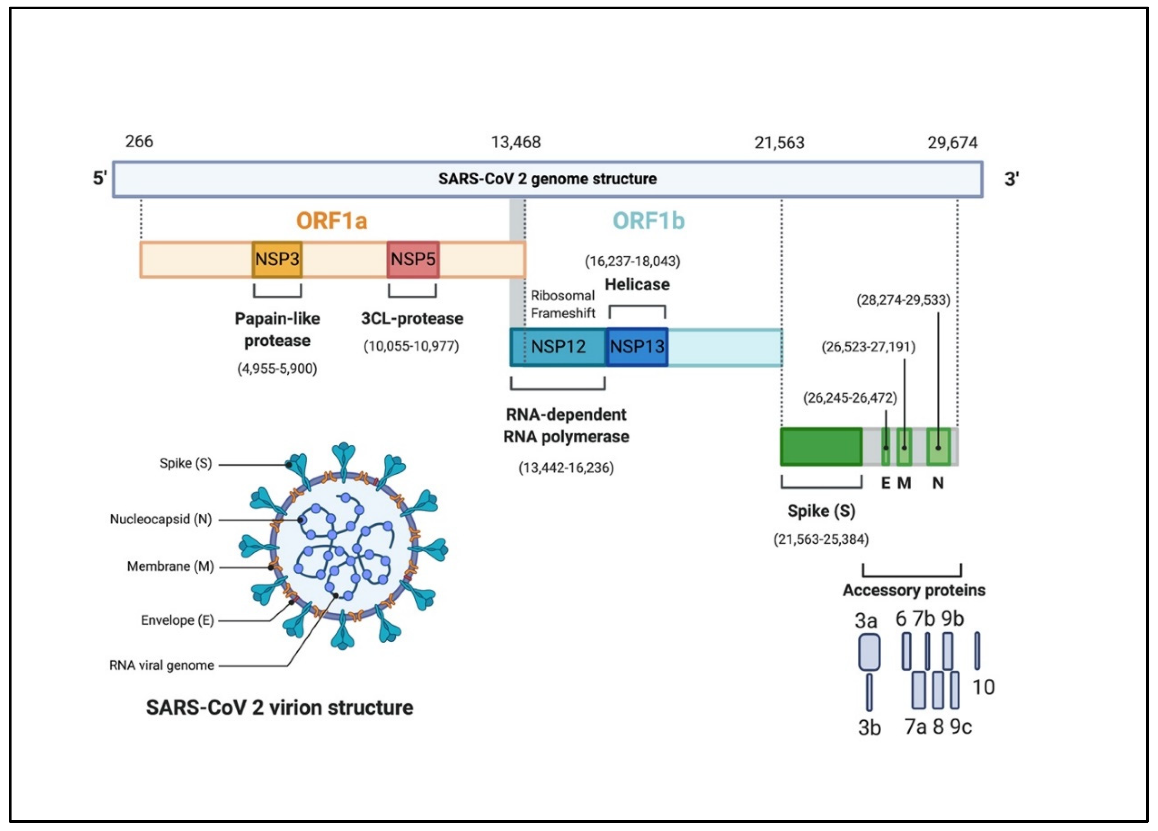
Each domain serves as a placeholder for genetic information that codes for several features of the virus. Overall, the genome structure of the virus contains at least six open reading frames (ORFs) (4). The 5’ domain makes up about two thirds of the genome. This region is comprised of the ORFs, ORF1a/b, which contain the information needed to code for the replication machinery of the virus (i.e. non-structural proteins) (1,3). The 3’ domain makes up the remaining third of the genome and codes for the structural proteins of the virus. Amongst these include the surface (S), envelope (E), membrane (M), and nucleocapsid (N) proteins. Additionally, the 3’ domain contains genetic information for accessory proteins which include ORF3a, ORF6, ORF7a, ORF7b, and ORF8 (3). Figure 2 illustrates the SARS-CoV-2 genome structure and its features.
Virus Particle Structure
The SARS-CoV-2 virion is spherical with a diameter of approximately 60-100 nm (2). Coronavirus particles contain four main structural proteins: the spike (S), membrane (M), envelope (E), and nucleocapsid (N) proteins. The individual functions of each structural protein can be described in the following manner:
- Spike (S)– recognizes and attaches to host cell receptors
- Membrane (M) – gives the virion its shape
- Envelope (E) – involved in assembly and release
- Nucleocapsid (N) – contains the RNA genome
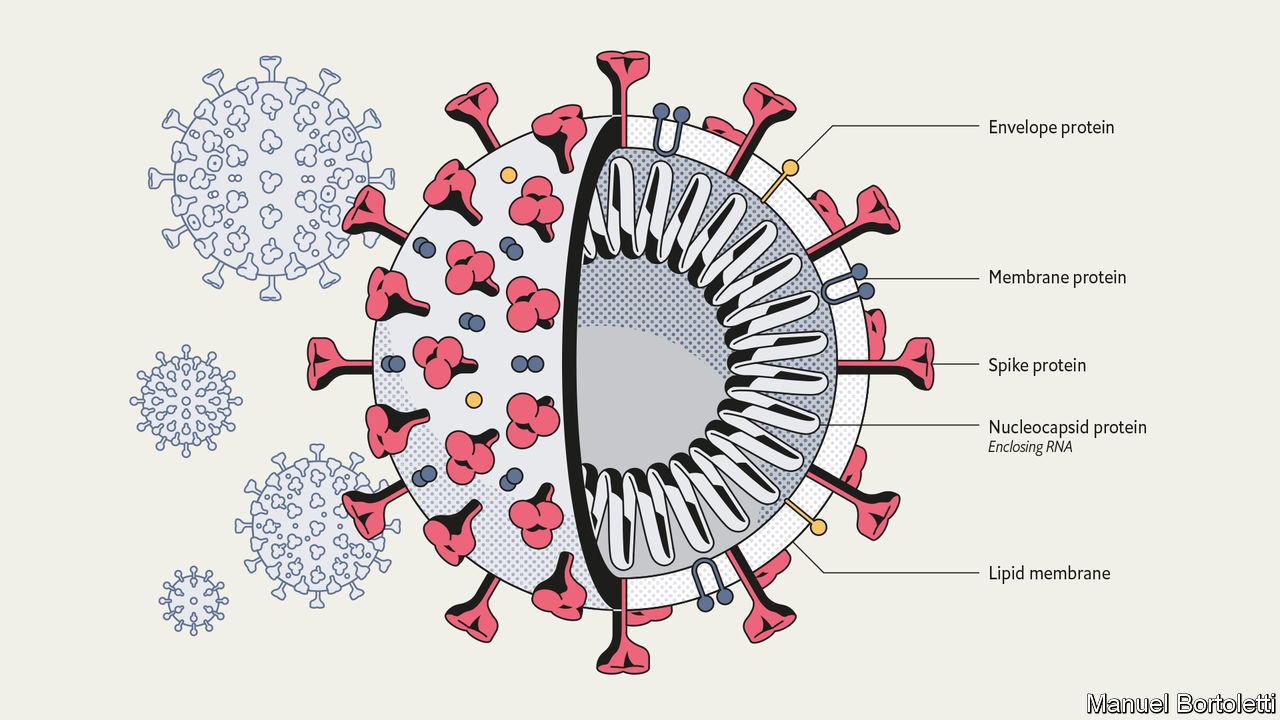
A prominent feature of the virus are the spike (S) proteins that surround the virion particle. These S proteins resemble the appearance of a solar corona, prompting the name coronavirus (1). Due to its ability to recognize the host cell and guide the mechanism of entry, the spike protein has been at the forefront of research as a possible site of action for vaccines and other pharmacological interventions.
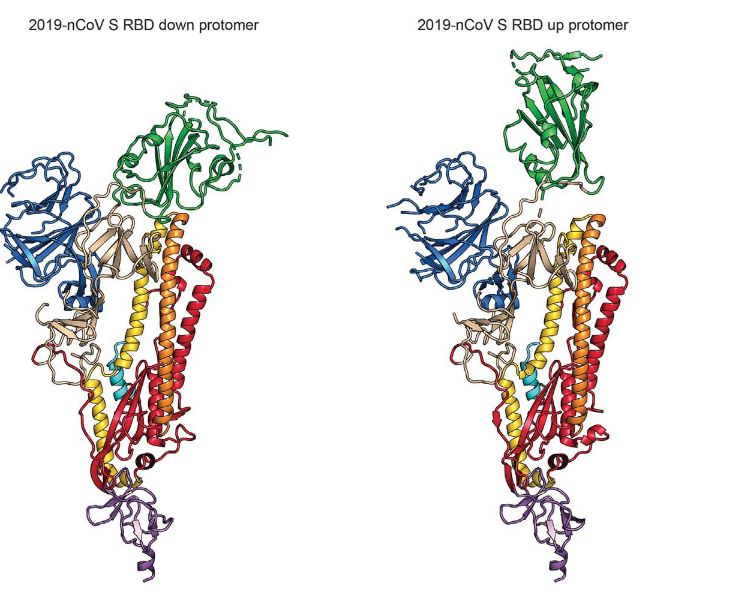
The S protein is a transmembrane glycoprotein composed of two subunits, S1 and S2, which are responsible for attachment and membrane fusion, respectively. This protein, emerging from the surface of the virion, exists in a homotrimer state (i.e. a protein which is composed of three identical polypeptide units) with a molecular weight of approximately 180 kDa (6). The S1 subunit is comprised of three receptor-binding heads which sit on top of a trimeric membrane fusion S2 stalk (7). Furthermore, Wrapp et al. explains that the S1 domain undergoes conformational movements that either hide or expose the domain in order to bind to the host cell receptor (the two states are referred to as the “down” conformation and the “up” conformation, respectively). Figure 4 shows a ribbon diagram of the S protein in these two states.
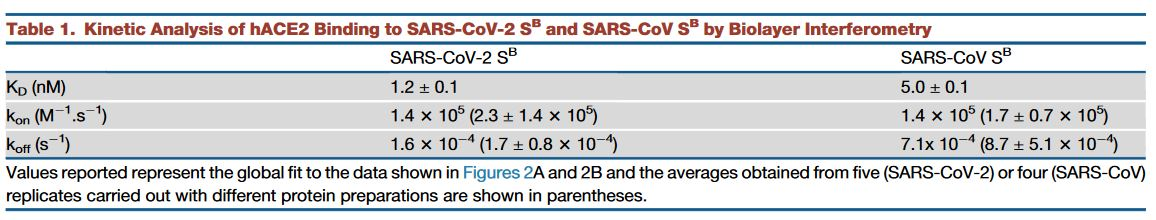
Various studies have shown that the S protein gains access to the host cell via the angiotensin-converting enzyme 2 (ACE2) receptor. Further investigations by Walls et al. have also seem to corroborate that SARS-CoV-2 spike protein recognizes the ACE2 receptor with a comparable affinity to 2002–2003 SARS-CoV isolates. Table 1 shows their enzyme kinetic analysis results.

Coronavirus Life Cycle
The primary site of infection of SARS-CoV-2 in humans are the bronchial epithelial cells, as well as type II pneumocytes, where the virus enters the cell via the angiotensin-converting enzyme 2 (ACE2) receptor. Moreover, ACE2 is also found in cells of other organs such as the heart, kidneys, and gastrointestinal tract which might further contribute to the virus’ pathogenicity. SARS-CoV-2 follows a similar pattern of infection to other coronaviruses. The life cycle of the virus, Figure 5, can be modeled in the following stages:
- Attachment and Entry
- Replicase Protein Expression
- Replication and Transcription
- Assembly and Release
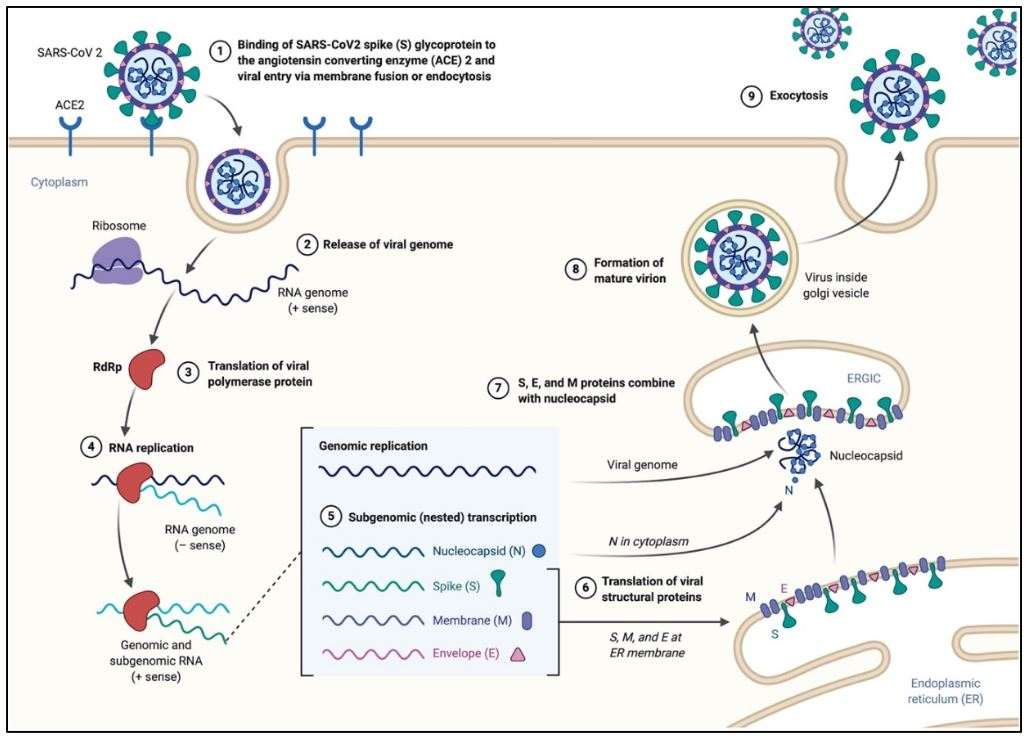
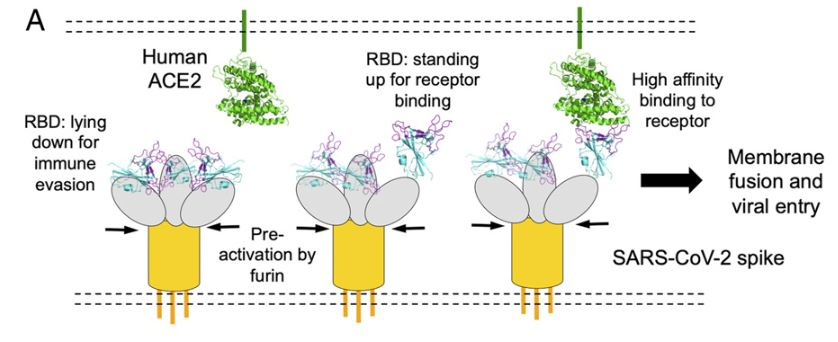
The process of cell entry begins with the binding of the S protein of the virus to the ACE2 receptor of the host cell. This initial binding occurs after the receptor-binding domain (RBD) of the S1 subunit of the S protein recognizes ACE2. Studies have indicated that the RBD continually switches between a “standing-up” position for receptor binding and a “lying-down” position for immune evasion (7). Next, membrane fusion occurs with the proteolytic activation of a furin cleavage site at the S1/S2 boundary of the spike protein by cell surface protease TMPRSS2 (cellular serine protease) and lysosomal proteases known as cathepsins (pH dependent cysteine proteases). This enzymatic action results in the dissociation of the RBD of the spike protein, S1, which exposes and causes a structural change of the fusion domain, S2, thus allowing the virus to enter the host cell (7).
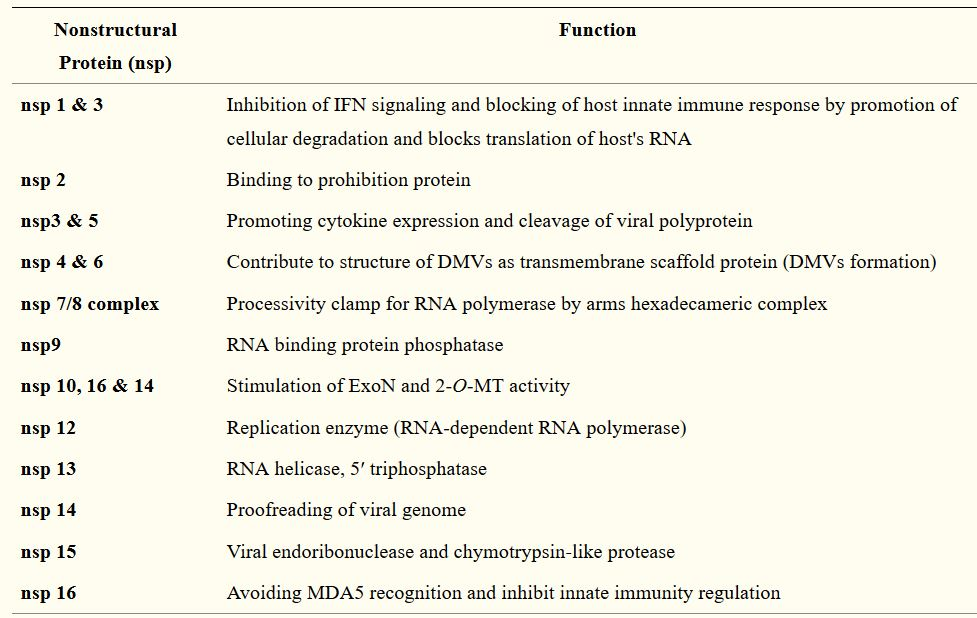
Following the entry of the virus into the host cell, the viral genome is released into the cell’s cytoplasm. The first gene segments to enter translation are those required to encode the replicase gene, which are nonstructural polyproteins involved in the genome replication (1,9). The replicase gene includes two ORFs, ORF1a and ORF1b, that encode two coterminal polyproteins, pp1a and pp1ab.1,9 Coronaviruses employ a mechanism of ribosomal frame shifting which utilizes a slippery sequence (5̘’-UUUAAAC-3̘’) and an RNA pseudoknot in order to express these polyproteins.1 Polyproteins pp1a and pp1ab are eventually cleaved into the nonstructural proteins (nsps) 1–11 and 1–16, respectively, by viral proteases that include the papain-like proteases (PLpro), encoded within nsp3, and a serine type protease Mpro, encoded by nsp5 (1,9). Table 2 describes the functions of the rest of the nsps.
Following the cleavage of the polyproteins into nsps, several of these proteins coalesce to assemble the replicase-transcriptase complex (RTC), which results in an environment suitable for RNA synthesis. Eventually, the assembly of the RTC is responsible for viral RNA replication and transcription of genomic and sub-genomic RNAs, the latter serving as the mRNAs responsible for the structural and accessory genes of the virus (1,9).During replication, the RTC produces an intermediate single-stranded negative-sense RNA (-ssRNA) intermediate that is used as a template for the full length single-stranded positive-sense RNA (+ssRNA) genome (1,4,9).
Following replication, sub-genomic RNAs are translated into the viral structural proteins (S, E, and M) and become inserted into the endoplasmic reticulum (ER) of the host cell. These structural proteins are then transported along the secretory pathway into the endoplasmic reticulum-Golgi intermediate compartment (ERGIC) where the viral genome, encapsidated by the N protein, buds with this newly created membrane along the ER to form the mature SARS-CoV-2 virions (1,9). These mature virions are transported in small vesicles and eventually exit the host cell via exocytosis.
Many questions still remain about SARS-CoV-2 and its pathogenicity in human beings. For example, it is still unclear why there is a significant difference in the severity of the respiratory syndrome caused by the virus, which has led to the current mortality rates, between different populations (particularly the elderly and those with chronic conditions). Continued efforts by the scientific community worldwide are vital. Research into the effects of the virus in a spectrum of hosts, as well as transmission patterns, is critical in order to develop effective strategies to treat those infected and prevent infection in the rest of the population.
References
- Fehr AR, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol. 2015;1282:1‐23. doi:10.1007/978-1-4939-2438-7_1
- Jin, Y.; Yang, H.; Ji, W.; Wu, W.; Chen, S.; Zhang, W.; Duan, G. Virology, Epidemiology, Pathogenesis, and Control of COVID-19. Viruses 2020, 12, 372.
- Khailany RA, Safdar M, Ozaslan M. Genomic characterization of a novel SARS-CoV-2 [published online ahead of print, 2020 Apr 16]. Gene Rep. 2020;19:100682. doi:10.1016/j.genrep.2020.100682
- Alanagreh L, Alzoughool F, Atoum M. The Human Coronavirus Disease COVID-19: Its Origin, Characteristics, and Insights into Potential Drugs and Its Mechanisms. Pathogens. 2020;9(5):E331. Published 2020 Apr 29. doi:10.3390/pathogens9050331
- Lu R, Zhao X, Li J, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565‐574. doi:10.1016/S0140-6736(20)30251-8
- Ou, X., Liu, Y., Lei, X. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11, 1620 (2020). https://doi.org/10.1038/s41467-020-15562-9
- Shang J, Wan Y, Luo C, et al. Cell entry mechanisms of SARS-CoV-2 [published online ahead of print, 2020 May 6]. Proc Natl Acad Sci U S A. 2020;202003138. doi:10.1073/pnas.2003138117
- Wrapp D, Wang N, Corbett KS, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260‐1263. doi:10.1126/science.abb2507
- Astuti I, Ysrafil. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response [published online ahead of print, 2020 Apr 18]. Diabetes Metab Syndr. 2020;14(4):407‐412. doi:10.1016/j.dsx.2020.04.020
- Yu WB, Tang GD, Zhang L, Corlett RT. Decoding the evolution and transmissions of the novel pneumonia coronavirus (SARS-CoV-2 / HCoV-19) using whole genomic data. Zool Res. 2020;41(3):247‐257. doi:10.24272/j.issn.2095-8137.2020.022
- Rehman SU, Shafique L, Ihsan A, Liu Q. Evolutionary Trajectory for the Emergence of Novel Coronavirus SARS-CoV-2. Pathogens. 2020;9(3):240. Published 2020 Mar 23. doi:10.3390/pathogens9030240
- American Society for Microbiology. Basic virology. (n.d.). ASM.org. https://asm.org/COVID/COVID-19-Research-Registry/Basic-Virology
- Corman VM, Muth D, Niemeyer D, Drosten C. Hosts and Sources of Endemic Human Coronaviruses. Adv Virus Res. 2018;100:163‐188. doi:10.1016/bs.aivir.2018.01.001